+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5td6 | ||||||
|---|---|---|---|---|---|---|---|
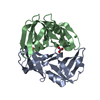
| Title | C. elegans FOG-3 BTG/Tob domain - H47N, C117A | ||||||
 Components Components | FOG-3 protein | ||||||
 Keywords Keywords | RNA BINDING PROTEIN / BTG/Tob / polymer / RNA binding | ||||||
| Function / homology |  Function and homology information Function and homology informationmasculinization of hermaphroditic germ-line / positive regulation of oocyte development / cell fate specification / nuclear periphery / transcription corepressor activity / regulation of gene expression / spermatogenesis / nucleus / cytoplasm Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.034 Å MOLECULAR REPLACEMENT / Resolution: 2.034 Å | ||||||
 Authors Authors | Aoki, S.T. / Bingman, C.A. / Wickens, M. / Kimble, J.E. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Cell Rep / Year: 2018 Journal: Cell Rep / Year: 2018Title: An RNA-Binding Multimer Specifies Nematode Sperm Fate. Authors: Aoki, S.T. / Porter, D.F. / Prasad, A. / Wickens, M. / Bingman, C.A. / Kimble, J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5td6.cif.gz 5td6.cif.gz | 123.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5td6.ent.gz pdb5td6.ent.gz | 95.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5td6.json.gz 5td6.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  5td6_validation.pdf.gz 5td6_validation.pdf.gz | 441.6 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  5td6_full_validation.pdf.gz 5td6_full_validation.pdf.gz | 443.3 KB | Display | |
| Data in XML |  5td6_validation.xml.gz 5td6_validation.xml.gz | 12.9 KB | Display | |
| Data in CIF |  5td6_validation.cif.gz 5td6_validation.cif.gz | 17.5 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/td/5td6 https://data.pdbj.org/pub/pdb/validation_reports/td/5td6 ftp://data.pdbj.org/pub/pdb/validation_reports/td/5td6 ftp://data.pdbj.org/pub/pdb/validation_reports/td/5td6 | HTTPS FTP |
-Related structure data
| Related structure data |  2d5rS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Symmetry | Helical symmetry: (Num. of operations: 1 / Rise per n subunits: 32.461 Å / Rotation per n subunits: 180 °) |
- Components
Components
| #1: Protein | Mass: 16216.328 Da / Num. of mol.: 2 / Fragment: UNP residues 1-137 / Mutation: H47N, C117A Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Chemical | ChemComp-SO4 / | #3: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.51 Å3/Da / Density % sol: 51.01 % |
|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, sitting drop / pH: 5.6 / Details: Mg Acetate, Na Citrate, PEG 10000 / PH range: 5.4-5.8 Temp details: set trays up at RT, moved to cold room (4 Celsius) |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 23-ID-D / Wavelength: 0.95372 Å / Beamline: 23-ID-D / Wavelength: 0.95372 Å |
| Detector | Type: DECTRIS PILATUS3 2M / Detector: PIXEL / Date: Jun 15, 2014 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.95372 Å / Relative weight: 1 |
| Reflection | Resolution: 2.034→29.22 Å / Num. obs: 21617 / % possible obs: 99.8 % / Redundancy: 6.8 % / CC1/2: 1 / Rmerge(I) obs: 0.0498 / Rsym value: 0.05399 / Net I/σ(I): 24.12 |
| Reflection shell | Resolution: 2.034→2.106 Å / Redundancy: 7 % / Rmerge(I) obs: 0.7383 / Mean I/σ(I) obs: 2.56 / CC1/2: 0.789 / % possible all: 98.18 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 2D5R Resolution: 2.034→29.22 Å / SU ML: 0.23 / Cross valid method: FREE R-VALUE / σ(F): 1.55 / Phase error: 21.9 Details: Author claims "Two densities were observed in the solvent-accessible area adjacent to residues 52-56 in both copies in the ASU. Density is observed at FoFc contour levels past 6 sigma. We ...Details: Author claims "Two densities were observed in the solvent-accessible area adjacent to residues 52-56 in both copies in the ASU. Density is observed at FoFc contour levels past 6 sigma. We attempted modeling of acetate (too small) and citrate (too large), both present in the crystallization conditions, but the fit was unsatisfactory. Thus, the final uploaded model does not account for these two large densities."
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.034→29.22 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 3.5469 Å / Origin y: 51.3325 Å / Origin z: 63.2053 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: all |
 Movie
Movie Controller
Controller












 PDBj
PDBj


