+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5m5q | ||||||
|---|---|---|---|---|---|---|---|
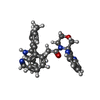
| Title | COPS5(2-257) IN COMPLEX WITH A AZAINDOLE (COMPOUND 4) | ||||||
 Components Components | COP9 signalosome complex subunit 5 | ||||||
 Keywords Keywords |  HYDROLASE / METAL PROTEASE / HYDROLASE / METAL PROTEASE /  COP9 SIGNALOSOME / COP9 SIGNALOSOME /  HYDROXYLASE HYDROXYLASE | ||||||
| Function / homology |  Function and homology information Function and homology information macrophage migration inhibitory factor binding / regulation of IRE1-mediated unfolded protein response / exosomal secretion / deNEDDylase activity / regulation of protein neddylation / eukaryotic translation initiation factor 3 complex / protein deneddylation / macrophage migration inhibitory factor binding / regulation of IRE1-mediated unfolded protein response / exosomal secretion / deNEDDylase activity / regulation of protein neddylation / eukaryotic translation initiation factor 3 complex / protein deneddylation /  COP9 signalosome / metal-dependent deubiquitinase activity / protein neddylation ... COP9 signalosome / metal-dependent deubiquitinase activity / protein neddylation ... macrophage migration inhibitory factor binding / regulation of IRE1-mediated unfolded protein response / exosomal secretion / deNEDDylase activity / regulation of protein neddylation / eukaryotic translation initiation factor 3 complex / protein deneddylation / macrophage migration inhibitory factor binding / regulation of IRE1-mediated unfolded protein response / exosomal secretion / deNEDDylase activity / regulation of protein neddylation / eukaryotic translation initiation factor 3 complex / protein deneddylation /  COP9 signalosome / metal-dependent deubiquitinase activity / protein neddylation / COP9 signalosome / metal-dependent deubiquitinase activity / protein neddylation /  Hydrolases; Acting on peptide bonds (peptidases) / protein deubiquitination / regulation of JNK cascade / Hydrolases; Acting on peptide bonds (peptidases) / protein deubiquitination / regulation of JNK cascade /  translation initiation factor activity / translation initiation factor activity /  post-translational protein modification / DNA Damage Recognition in GG-NER / Formation of TC-NER Pre-Incision Complex / positive regulation of DNA-binding transcription factor activity / post-translational protein modification / DNA Damage Recognition in GG-NER / Formation of TC-NER Pre-Incision Complex / positive regulation of DNA-binding transcription factor activity /  metallopeptidase activity / metallopeptidase activity /  synaptic vesicle / Cargo recognition for clathrin-mediated endocytosis / synaptic vesicle / Cargo recognition for clathrin-mediated endocytosis /  Neddylation / Neddylation /  transcription coactivator activity / transcription coactivator activity /  regulation of cell cycle / regulation of cell cycle /  translation / translation /  chromatin / negative regulation of apoptotic process / perinuclear region of cytoplasm / chromatin / negative regulation of apoptotic process / perinuclear region of cytoplasm /  enzyme binding / positive regulation of transcription by RNA polymerase II / enzyme binding / positive regulation of transcription by RNA polymerase II /  proteolysis / proteolysis /  nucleoplasm / nucleoplasm /  metal ion binding / metal ion binding /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.2 Å MOLECULAR REPLACEMENT / Resolution: 2.2 Å | ||||||
 Authors Authors | Renatus, M. / Altmann, E. | ||||||
 Citation Citation |  Journal: Angew. Chem. Int. Ed. Engl. / Year: 2017 Journal: Angew. Chem. Int. Ed. Engl. / Year: 2017Title: Azaindoles as Zinc-Binding Small-Molecule Inhibitors of the JAMM Protease CSN5. Authors: Altmann, E. / Erbel, P. / Renatus, M. / Schaefer, M. / Schlierf, A. / Druet, A. / Kieffer, L. / Sorge, M. / Pfister, K. / Hassiepen, U. / Jones, M. / Ruedisser, S. / Ostermeier, D. / ...Authors: Altmann, E. / Erbel, P. / Renatus, M. / Schaefer, M. / Schlierf, A. / Druet, A. / Kieffer, L. / Sorge, M. / Pfister, K. / Hassiepen, U. / Jones, M. / Ruedisser, S. / Ostermeier, D. / Martoglio, B. / Jefferson, A.B. / Quancard, J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5m5q.cif.gz 5m5q.cif.gz | 62.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5m5q.ent.gz pdb5m5q.ent.gz | 43.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5m5q.json.gz 5m5q.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/m5/5m5q https://data.pdbj.org/pub/pdb/validation_reports/m5/5m5q ftp://data.pdbj.org/pub/pdb/validation_reports/m5/5m5q ftp://data.pdbj.org/pub/pdb/validation_reports/m5/5m5q | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein |  / Signalosome subunit 5 / Jun activation domain-binding protein 1 / Signalosome subunit 5 / Jun activation domain-binding protein 1Mass: 28950.980 Da / Num. of mol.: 1 / Fragment: UNP residues 2-257 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: COPS5, CSN5, JAB1 / Production host: Homo sapiens (human) / Gene: COPS5, CSN5, JAB1 / Production host:   Escherichia coli (E. coli) Escherichia coli (E. coli)References: UniProt: Q92905,  Hydrolases; Acting on peptide bonds (peptidases) Hydrolases; Acting on peptide bonds (peptidases) |
|---|---|
| #2: Chemical | ChemComp-ZN / |
| #3: Chemical | ChemComp-7K1 / |
| #4: Chemical | ChemComp-MPD / ( 2-Methyl-2,4-pentanediol 2-Methyl-2,4-pentanediol |
| #5: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.12 Å3/Da / Density % sol: 60.63 % |
|---|---|
Crystal grow | Temperature: 277 K / Method: vapor diffusion, sitting drop / pH: 6 / Details: 35% MPD, 0.2M LiSO4, 0,1 M Mes pH 6.0 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SLS SLS  / Beamline: X10SA / Wavelength: 1 Å / Beamline: X10SA / Wavelength: 1 Å |
| Detector | Type: DECTRIS PILATUS 2M / Detector: PIXEL / Date: Aug 22, 2012 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1 Å / Relative weight: 1 : 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→100 Å / Num. obs: 18226 / % possible obs: 100 % / Redundancy: 6.8 % / Biso Wilson estimate: 57.66 Å2 / Rsym value: 0.045 / Net I/σ(I): 22.5 |
| Reflection shell | Resolution: 2.2→2.26 Å / Redundancy: 6.7 % / Mean I/σ(I) obs: 2.8 / % possible all: 99.9 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: in house model Resolution: 2.2→21.73 Å / Cor.coef. Fo:Fc: 0.937 / Cor.coef. Fo:Fc free: 0.93 / SU R Cruickshank DPI: 0.178 / Cross valid method: THROUGHOUT / σ(F): 0 / SU R Blow DPI: 0.173 / SU Rfree Blow DPI: 0.149 / SU Rfree Cruickshank DPI: 0.153
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 68.84 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→21.73 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.2→2.33 Å / Rfactor Rfree error: 0 / Total num. of bins used: 9
|
 Movie
Movie Controller
Controller











 PDBj
PDBj