| Entry | Database: PDB / ID: 5l71
|
|---|
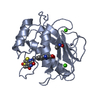
| Title | Crystal structure of mouse phospholipid hydroperoxide glutathione peroxidase 4 (GPx4) |
|---|
 Components Components | Phospholipid hydroperoxide glutathione peroxidase, mitochondrial |
|---|
 Keywords Keywords | OXIDOREDUCTASE / phospholipid hydroperoxide glutathione peroxidase 4 (GPx4) / selenocysteine |
|---|
| Function / homology |  Function and homology information Function and homology information
Synthesis of 12-eicosatetraenoic acid derivatives / Biosynthesis of D-series resolvins / Biosynthesis of E-series 18(S)-resolvins / Biosynthesis of aspirin-triggered D-series resolvins / Biosynthesis of E-series 18(R)-resolvins / phospholipid-hydroperoxide glutathione peroxidase / phospholipid-hydroperoxide glutathione peroxidase activity / selenium binding / Synthesis of 15-eicosatetraenoic acid derivatives / glutathione peroxidase ...Synthesis of 12-eicosatetraenoic acid derivatives / Biosynthesis of D-series resolvins / Biosynthesis of E-series 18(S)-resolvins / Biosynthesis of aspirin-triggered D-series resolvins / Biosynthesis of E-series 18(R)-resolvins / phospholipid-hydroperoxide glutathione peroxidase / phospholipid-hydroperoxide glutathione peroxidase activity / selenium binding / Synthesis of 15-eicosatetraenoic acid derivatives / glutathione peroxidase / lipoxygenase pathway / arachidonate metabolic process / glutathione peroxidase activity / dendrite development / negative regulation of ferroptosis / protein polymerization / cerebellum development / multicellular organism growth / response to estradiol / nuclear envelope / chromatin organization / response to oxidative stress / spermatogenesis / response to lipopolysaccharide / mitochondrial inner membrane / apoptotic process / protein-containing complex / mitochondrion / identical protein binding / nucleus / cytosolSimilarity search - Function Glutathione peroxidase active site / Glutathione peroxidases active site. / Glutathione peroxidase / Glutathione peroxidase conserved site / Glutathione peroxidase / Glutathione peroxidases signature 2. / Glutathione peroxidase profile. / Glutaredoxin / Glutaredoxin / Thioredoxin-like superfamily ...Glutathione peroxidase active site / Glutathione peroxidases active site. / Glutathione peroxidase / Glutathione peroxidase conserved site / Glutathione peroxidase / Glutathione peroxidases signature 2. / Glutathione peroxidase profile. / Glutaredoxin / Glutaredoxin / Thioredoxin-like superfamily / 3-Layer(aba) Sandwich / Alpha BetaSimilarity search - Domain/homology |
|---|
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å |
|---|
 Authors Authors | Janowski, R. / Scanu, S. / Madl, T. / Niessing, D. |
|---|
 Citation Citation |  Journal: Acta Crystallogr.,Sect.F / Year: 2016 Journal: Acta Crystallogr.,Sect.F / Year: 2016
Title: Crystal and solution structural studies of mouse phospholipid hydroperoxide glutathione peroxidase 4.
Authors: Janowski, R. / Scanu, S. / Niessing, D. / Madl, T. |
|---|
| History | | Deposition | Jun 1, 2016 | Deposition site: PDBE / Processing site: PDBE |
|---|
| Revision 1.0 | Oct 19, 2016 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Jan 10, 2024 | Group: Data collection / Database references / Refinement description
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession |
|---|
|
|---|
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å
MOLECULAR REPLACEMENT / Resolution: 1.8 Å  Authors
Authors Citation
Citation Journal: Acta Crystallogr.,Sect.F / Year: 2016
Journal: Acta Crystallogr.,Sect.F / Year: 2016 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5l71.cif.gz
5l71.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5l71.ent.gz
pdb5l71.ent.gz PDB format
PDB format 5l71.json.gz
5l71.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/l7/5l71
https://data.pdbj.org/pub/pdb/validation_reports/l7/5l71 ftp://data.pdbj.org/pub/pdb/validation_reports/l7/5l71
ftp://data.pdbj.org/pub/pdb/validation_reports/l7/5l71
 Links
Links Assembly
Assembly
 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  SLS
SLS  / Beamline: X06SA / Wavelength: 0.99999 Å
/ Beamline: X06SA / Wavelength: 0.99999 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller











 PDBj
PDBj



