| Entry | Database: PDB / ID: 5f67
|
|---|
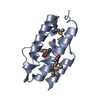
| Title | An exquisitely specific PDZ/target recognition revealed by the structure of INAD PDZ3 in complex with TRP channel tail |
|---|
 Components Components | - Inactivation-no-after-potential D protein
- TRP C terminal Tail
|
|---|
 Keywords Keywords | PROTEIN BINDING / INAD / TRP / PDZ / atypical |
|---|
| Function / homology |  Function and homology information Function and homology information
phospholipase C-activating opsin-mediated signaling pathway / Ion homeostasis / TRP channels / myosin III binding / rhabdomere microvillus membrane / Antigen activates B Cell Receptor (BCR) leading to generation of second messengers / detection of light stimulus involved in sensory perception / inaD signaling complex / cellular response to anoxia / negative regulation of opsin-mediated signaling pathway ...phospholipase C-activating opsin-mediated signaling pathway / Ion homeostasis / TRP channels / myosin III binding / rhabdomere microvillus membrane / Antigen activates B Cell Receptor (BCR) leading to generation of second messengers / detection of light stimulus involved in sensory perception / inaD signaling complex / cellular response to anoxia / negative regulation of opsin-mediated signaling pathway / Antigen processing: Ubiquitination & Proteasome degradation / rhabdomere / cellular response to light stimulus / store-operated calcium channel activity / detection of light stimulus involved in visual perception / olfactory learning / inositol 1,4,5 trisphosphate binding / cation channel complex / retina homeostasis / : / myosin binding / phototransduction, visible light / photoreceptor activity / phototransduction / response to light stimulus / regulation of cytosolic calcium ion concentration / visual perception / mitochondrion organization / sensory perception of sound / calcium ion transmembrane transport / calcium channel activity / sensory perception of smell / calcium ion transport / intracellular protein localization / signaling receptor complex adaptor activity / calmodulin binding / protein heterodimerization activity / protein homodimerization activity / membrane / identical protein binding / plasma membraneSimilarity search - Function : / Transient receptor ion channel domain / Transient receptor ion channel II / Transient receptor ion channel II / Transient receptor potential channel, canonical / PDZ domain / Pdz3 Domain / Ankyrin repeat / PDZ domain / PDZ domain profile. ...: / Transient receptor ion channel domain / Transient receptor ion channel II / Transient receptor ion channel II / Transient receptor potential channel, canonical / PDZ domain / Pdz3 Domain / Ankyrin repeat / PDZ domain / PDZ domain profile. / Domain present in PSD-95, Dlg, and ZO-1/2. / PDZ domain / PDZ superfamily / Ankyrin repeat profile. / Ankyrin repeats (3 copies) / Ankyrin repeat region circular profile. / ankyrin repeats / Ankyrin repeat / Ankyrin repeat-containing domain superfamily / Ion transport domain / Ion transport protein / Roll / Mainly BetaSimilarity search - Domain/homology |
|---|
| Biological species |   Drosophila melanogaster (fruit fly) Drosophila melanogaster (fruit fly) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.76 Å MOLECULAR REPLACEMENT / Resolution: 1.76 Å |
|---|
 Authors Authors | Ye, F. / Shang, Y. / Liu, W. / Zhang, M. |
|---|
 Citation Citation |  Journal: Structure / Year: 2016 Journal: Structure / Year: 2016
Title: An Exquisitely Specific PDZ/Target Recognition Revealed by the Structure of INAD PDZ3 in Complex with TRP Channel Tail
Authors: Ye, F. / Liu, W. / Shang, Y. / Zhang, M. |
|---|
| History | | Deposition | Dec 5, 2015 | Deposition site: RCSB / Processing site: PDBJ |
|---|
| Revision 1.0 | Feb 24, 2016 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Mar 16, 2016 | Group: Database references |
|---|
| Revision 1.2 | Nov 8, 2023 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Refinement description
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / citation / database_2 / pdbx_initial_refinement_model / pdbx_struct_oper_list
Item: _citation.journal_id_CSD / _database_2.pdbx_DOI ..._citation.journal_id_CSD / _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_struct_oper_list.symmetry_operation |
|---|
|
|---|
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.76 Å
MOLECULAR REPLACEMENT / Resolution: 1.76 Å  Authors
Authors Citation
Citation Journal: Structure / Year: 2016
Journal: Structure / Year: 2016 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5f67.cif.gz
5f67.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5f67.ent.gz
pdb5f67.ent.gz PDB format
PDB format 5f67.json.gz
5f67.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/f6/5f67
https://data.pdbj.org/pub/pdb/validation_reports/f6/5f67 ftp://data.pdbj.org/pub/pdb/validation_reports/f6/5f67
ftp://data.pdbj.org/pub/pdb/validation_reports/f6/5f67
 Links
Links Assembly
Assembly


 Components
Components


 X-RAY DIFFRACTION
X-RAY DIFFRACTION Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  SSRF
SSRF  / Beamline: BL17U / Wavelength: 0.97923 Å
/ Beamline: BL17U / Wavelength: 0.97923 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller











 PDBj
PDBj



