[English] 日本語
 Yorodumi
Yorodumi- PDB-4tsm: MBP-fusion protein of PilA1 from C. difficile R20291 residues 26-166 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4tsm | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
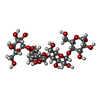
| Title | MBP-fusion protein of PilA1 from C. difficile R20291 residues 26-166 | |||||||||
 Components Components | maltose-binding protein, pilin chimera | |||||||||
 Keywords Keywords | CELL ADHESION / Pilin / T4P / fusion | |||||||||
| Function / homology |  Function and homology information Function and homology informationprotein secretion by the type II secretion system / type II protein secretion system complex / detection of maltose stimulus / maltose transport complex / carbohydrate transport / membrane => GO:0016020 / carbohydrate transmembrane transporter activity / maltose binding / maltose transport / maltodextrin transmembrane transport ...protein secretion by the type II secretion system / type II protein secretion system complex / detection of maltose stimulus / maltose transport complex / carbohydrate transport / membrane => GO:0016020 / carbohydrate transmembrane transporter activity / maltose binding / maltose transport / maltodextrin transmembrane transport / ATP-binding cassette (ABC) transporter complex, substrate-binding subunit-containing / ATP-binding cassette (ABC) transporter complex / cell chemotaxis / outer membrane-bounded periplasmic space / periplasmic space / DNA damage response / membrane Similarity search - Function | |||||||||
| Biological species |   Peptoclostridium difficile R20291 (bacteria) Peptoclostridium difficile R20291 (bacteria) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.899 Å MOLECULAR REPLACEMENT / Resolution: 1.899 Å | |||||||||
 Authors Authors | Piepenbrink, K.H. / Sundberg, E.J. | |||||||||
| Funding support |  United States, 2items United States, 2items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2015 Journal: Structure / Year: 2015Title: Structural and Evolutionary Analyses Show Unique Stabilization Strategies in the Type IV Pili of Clostridium difficile. Authors: Piepenbrink, K.H. / Maldarelli, G.A. / Martinez de la Pena, C.F. / Dingle, T.C. / Mulvey, G.L. / Lee, A. / von Rosenvinge, E. / Armstrong, G.D. / Donnenberg, M.S. / Sundberg, E.J. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4tsm.cif.gz 4tsm.cif.gz | 319.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4tsm.ent.gz pdb4tsm.ent.gz | 258 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4tsm.json.gz 4tsm.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ts/4tsm https://data.pdbj.org/pub/pdb/validation_reports/ts/4tsm ftp://data.pdbj.org/pub/pdb/validation_reports/ts/4tsm ftp://data.pdbj.org/pub/pdb/validation_reports/ts/4tsm | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4ogmSC  4pe2C C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 56380.512 Da / Num. of mol.: 3 Fragment: UNP residues 27-392 (MBP), UNP residues 35-173 (PilA1) Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Peptoclostridium difficile R20291 (bacteria) Peptoclostridium difficile R20291 (bacteria)Gene: malE, HMPREF9530_03068, CDR20291_3350 / Production host:  References: UniProt: D8A942, UniProt: C9YRY2, UniProt: P0AEX9*PLUS #2: Polysaccharide | #3: Chemical | ChemComp-TMO / #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.85 Å3/Da / Density % sol: 56.81 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 8.5 Details: 20% PEG2000 MME, 0.2 M trimethylamine N-oxide, 0.1 M Tris-HCl, pH 8.5 |
-Data collection
| Diffraction | Mean temperature: 99.1 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL11-1 / Wavelength: 0.97947 Å / Beamline: BL11-1 / Wavelength: 0.97947 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Jun 10, 2014 |
| Radiation | Monochromator: Si(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.97947 Å / Relative weight: 1 |
| Reflection | Resolution: 1.899→38.58 Å / Num. all: 146602 / Num. obs: 139534 / % possible obs: 94.9 % / Redundancy: 7.1 % / Rmerge(I) obs: 0.1776 / Net I/av σ(I): 8.8 / Net I/σ(I): 1.89 |
| Reflection shell | Resolution: 1.899→1.966 Å / Redundancy: 7 % / Rmerge(I) obs: 1.844 / Mean I/σ(I) obs: 1.14 / % possible all: 92.48 |
- Processing
Processing
| Software |
| ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB entry 4OGM Resolution: 1.899→38.58 Å / Cross valid method: FREE R-VALUE
| ||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.899→38.58 Å
| ||||||||||||||
| LS refinement shell | Resolution: 1.899→1.966 Å / Rfactor Rfree: 0.3136 / Rfactor Rwork: 0.2887 |
 Movie
Movie Controller
Controller












 PDBj
PDBj