+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4ciu | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of E. coli ClpB | ||||||
 Components Components | CHAPERONE PROTEIN CLPB | ||||||
 Keywords Keywords | CHAPERONE / AAA+ / ATPASE | ||||||
| Function / homology |  Function and homology information Function and homology informationcellular response to heat / response to heat / protein refolding / ATP hydrolysis activity / ATP binding / membrane / identical protein binding / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 3.5 Å SAD / Resolution: 3.5 Å | ||||||
 Authors Authors | Kopp, J. / Sinning, I. / Bukau, B. / Kummer, E. / Mogk, A. | ||||||
 Citation Citation |  Journal: Elife / Year: 2014 Journal: Elife / Year: 2014Title: Head-to-tail interactions of the coiled-coil domains regulate ClpB activity and cooperation with Hsp70 in protein disaggregation. Authors: Marta Carroni / Eva Kummer / Yuki Oguchi / Petra Wendler / Daniel K Clare / Irmgard Sinning / Jürgen Kopp / Axel Mogk / Bernd Bukau / Helen R Saibil /   Abstract: The hexameric AAA+ chaperone ClpB reactivates aggregated proteins in cooperation with the Hsp70 system. Essential for disaggregation, the ClpB middle domain (MD) is a coiled-coil propeller that binds ...The hexameric AAA+ chaperone ClpB reactivates aggregated proteins in cooperation with the Hsp70 system. Essential for disaggregation, the ClpB middle domain (MD) is a coiled-coil propeller that binds Hsp70. Although the ClpB subunit structure is known, positioning of the MD in the hexamer and its mechanism of action are unclear. We obtained electron microscopy (EM) structures of the BAP variant of ClpB that binds the protease ClpP, clearly revealing MD density on the surface of the ClpB ring. Mutant analysis and asymmetric reconstructions show that MDs adopt diverse positions in a single ClpB hexamer. Adjacent, horizontally oriented MDs form head-to-tail contacts and repress ClpB activity by preventing Hsp70 interaction. Tilting of the MD breaks this contact, allowing Hsp70 binding, and releasing the contact in adjacent subunits. Our data suggest a wavelike activation of ClpB subunits around the ring.DOI: http://dx.doi.org/10.7554/eLife.02481.001. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4ciu.cif.gz 4ciu.cif.gz | 283.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4ciu.ent.gz pdb4ciu.ent.gz | 230.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4ciu.json.gz 4ciu.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ci/4ciu https://data.pdbj.org/pub/pdb/validation_reports/ci/4ciu ftp://data.pdbj.org/pub/pdb/validation_reports/ci/4ciu ftp://data.pdbj.org/pub/pdb/validation_reports/ci/4ciu | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2555C  2556C  2557C  2558C 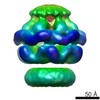 2559C 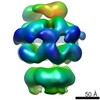 2560C  2561C  2562C  2563C  4d2qC  4d2uC  4d2xC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 82834.430 Da / Num. of mol.: 1 / Fragment: RESIDUES 143-857 / Mutation: YES Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Production host:  Variant (production host): DELTACLPB / References: UniProt: P63284 | ||
|---|---|---|---|
| #2: Chemical | | Has protein modification | Y | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.39 Å3/Da / Density % sol: 61 % / Description: NONE |
|---|---|
| Crystal grow | Temperature: 291 K Details: 1.5 M AMSO4, 0.4 % PEG 400, 0.1 M HEPES PH 7.5, 22 % GLYCEROL AT 291 K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID23-1 / Wavelength: 0.9794 / Beamline: ID23-1 / Wavelength: 0.9794 |
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Nov 7, 2011 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9794 Å / Relative weight: 1 |
| Reflection | Resolution: 3.5→81.2 Å / Num. obs: 14024 / % possible obs: 99.9 % / Observed criterion σ(I): 1 / Redundancy: 14.5 % / Biso Wilson estimate: 121 Å2 / Rmerge(I) obs: 0.09 / Net I/σ(I): 16.8 |
| Reflection shell | Resolution: 3.5→3.69 Å / Redundancy: 14.8 % / Rmerge(I) obs: 0.53 / Mean I/σ(I) obs: 5.4 / % possible all: 100 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  SAD SADStarting model: NONE Resolution: 3.5→36.8 Å / SU ML: 0.4 / σ(F): 1.37 / Phase error: 28.05 / Stereochemistry target values: MLHL
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL PHENIX 1.8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 91.1 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 3.5→36.8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj





