[English] 日本語
 Yorodumi
Yorodumi- PDB-3zzy: Crystal structure of a Raver1 PRI3 peptide in complex with polypy... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3zzy | ||||||
|---|---|---|---|---|---|---|---|
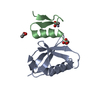
| Title | Crystal structure of a Raver1 PRI3 peptide in complex with polypyrimidine tract binding protein RRM2 | ||||||
 Components Components |
| ||||||
 Keywords Keywords | PROTEIN BINDING / PEPTIDE BINDING / RNA RECOGNITION MOTIF | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of muscle cell differentiation / poly-pyrimidine tract binding / IRES-dependent viral translational initiation / positive regulation of calcineurin-NFAT signaling cascade / pre-mRNA binding / FGFR2 alternative splicing / negative regulation of RNA splicing / negative regulation of neuron differentiation / regulation of alternative mRNA splicing, via spliceosome / regulation of RNA splicing ...negative regulation of muscle cell differentiation / poly-pyrimidine tract binding / IRES-dependent viral translational initiation / positive regulation of calcineurin-NFAT signaling cascade / pre-mRNA binding / FGFR2 alternative splicing / negative regulation of RNA splicing / negative regulation of neuron differentiation / regulation of alternative mRNA splicing, via spliceosome / regulation of RNA splicing / negative regulation of mRNA splicing, via spliceosome / regulation of cell differentiation / Processing of Capped Intron-Containing Pre-mRNA / mRNA Splicing - Major Pathway / neurogenesis / RNA splicing / mRNA processing / mRNA binding / nucleolus / positive regulation of transcription by RNA polymerase II / RNA binding / extracellular exosome / nucleoplasm / membrane / nucleus / cytoplasm Similarity search - Function | ||||||
| Biological species |  HOMO SAPIENS (human) HOMO SAPIENS (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.4 Å MOLECULAR REPLACEMENT / Resolution: 1.4 Å | ||||||
 Authors Authors | Joshi, A. / Kotik-Kogan, O. / Curry, S. | ||||||
 Citation Citation |  Journal: Structure / Year: 2011 Journal: Structure / Year: 2011Title: Crystallographic Analysis of Polypyrimidine Tract-Binding Protein-Raver1 Interactions Involved in Regulation of Alternative Splicing. Authors: Joshi, A. / Coelho, M.B. / Kotik-Kogan, O. / Simpson, P.J. / Matthews, S.J. / Smith, C.W. / Curry, S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3zzy.cif.gz 3zzy.cif.gz | 61.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3zzy.ent.gz pdb3zzy.ent.gz | 44.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3zzy.json.gz 3zzy.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/zz/3zzy https://data.pdbj.org/pub/pdb/validation_reports/zz/3zzy ftp://data.pdbj.org/pub/pdb/validation_reports/zz/3zzy ftp://data.pdbj.org/pub/pdb/validation_reports/zz/3zzy | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3zzzC  1sjrS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (-0.2051, 0.0424, 0.9778), Vector: |
- Components
Components
| #1: Protein | Mass: 14169.072 Da / Num. of mol.: 2 / Fragment: RNA RECOGNITION MOTIF 2, RESIDUES 172-301 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  HOMO SAPIENS (human) / Production host: HOMO SAPIENS (human) / Production host:  #2: Protein/peptide | Mass: 1467.710 Da / Num. of mol.: 2 / Fragment: MOTIF PRI3, RESIDUES 496-507 / Mutation: YES Source method: isolated from a genetically manipulated source Details: THIS FRAGMENT IS ACTUALLY FUSED TO MOLECULE 1, SEE REMARK 999 Source: (gene. exp.)   #3: Water | ChemComp-HOH / | Has protein modification | Y | Sequence details | CHAIN C AND D FIRST 4 RESIDUES (GAMG) ARE VECTOR DERIVED. RESIDUES 5 - 16 CORRESPOND TO RAVER1 PRI3 ...CHAIN C AND D FIRST 4 RESIDUES (GAMG) ARE VECTOR DERIVED. RESIDUES 5 - 16 CORRESPOND | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.01 Å3/Da / Density % sol: 38.94 % / Description: NONE |
|---|---|
| Crystal grow | pH: 6.5 / Details: SEE PAPER., pH 6.5 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  EMBL/DESY, HAMBURG EMBL/DESY, HAMBURG  / Beamline: X11 / Wavelength: 0.8123 / Beamline: X11 / Wavelength: 0.8123 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Dec 15, 2008 / Details: MIRROR |
| Radiation | Monochromator: GEMANIUM CRYSTAL / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.8123 Å / Relative weight: 1 |
| Reflection | Resolution: 1.4→30.32 Å / Num. obs: 50129 / % possible obs: 98.9 % / Observed criterion σ(I): 0 / Redundancy: 5.6 % / Biso Wilson estimate: 19.5 Å2 / Rmerge(I) obs: 0.06 / Net I/σ(I): 15.4 |
| Reflection shell | Resolution: 1.4→1.48 Å / Redundancy: 5.3 % / Rmerge(I) obs: 0.35 / Mean I/σ(I) obs: 5.3 / % possible all: 96.1 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1SJR Resolution: 1.4→29.01 Å / Rfactor Rfree error: 0.005 / Data cutoff high absF: 839936.87 / Data cutoff low absF: 0 / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: MLF Details: BULK SOLVENT MODEL USED. THE CRYSTALLISED PROTEIN HAS A RAVER1 PEPTIDE GENETICALLY FUSED AS AN N-TERMINAL EXTENSION OF PTB RRM2. OWING TO LINKER DISORDER WE CANNOT DETERMINE WHICH RRM DOMAIN ...Details: BULK SOLVENT MODEL USED. THE CRYSTALLISED PROTEIN HAS A RAVER1 PEPTIDE GENETICALLY FUSED AS AN N-TERMINAL EXTENSION OF PTB RRM2. OWING TO LINKER DISORDER WE CANNOT DETERMINE WHICH RRM DOMAIN (CHAINS A, B) IS COVALENTLY LINKED TO WHICH RAVER1 PEPTIDE (CHAINS C,D).
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 63.17 Å2 / ksol: 0.45 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 22.1 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.4→29.01 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | NCS model details: NONE | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.4→1.49 Å / Rfactor Rfree error: 0.015 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller


















 PDBj
PDBj


