+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3ygs | ||||||
|---|---|---|---|---|---|---|---|
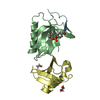
| Title | APAF-1 CARD IN COMPLEX WITH PRODOMAIN OF PROCASPASE-9 | ||||||
 Components Components |
| ||||||
 Keywords Keywords | APOPTOSIS / CASPASE ACTIVATION / CASPASE RECRUITMENT / RECOGNITION COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationcaspase-9 / response to G1 DNA damage checkpoint signaling / caspase complex / Formation of apoptosome / regulation of apoptotic DNA fragmentation / apoptosome / activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c / leukocyte apoptotic process / glial cell apoptotic process / response to cobalt ion ...caspase-9 / response to G1 DNA damage checkpoint signaling / caspase complex / Formation of apoptosome / regulation of apoptotic DNA fragmentation / apoptosome / activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c / leukocyte apoptotic process / glial cell apoptotic process / response to cobalt ion / cysteine-type endopeptidase activity involved in apoptotic signaling pathway / Caspase activation via Dependence Receptors in the absence of ligand / Activation of caspases through apoptosome-mediated cleavage / Regulation of the apoptosome activity / cysteine-type endopeptidase activity involved in apoptotic process / SMAC (DIABLO) binds to IAPs / SMAC(DIABLO)-mediated dissociation of IAP:caspase complexes / AKT phosphorylates targets in the cytosol / fibroblast apoptotic process / epithelial cell apoptotic process / platelet formation / TP53 Regulates Transcription of Caspase Activators and Caspases / Transcriptional Regulation by E2F6 / Constitutive Signaling by AKT1 E17K in Cancer / cysteine-type endopeptidase activator activity involved in apoptotic process / protein maturation / intrinsic apoptotic signaling pathway in response to endoplasmic reticulum stress / signal transduction in response to DNA damage / forebrain development / cardiac muscle cell apoptotic process / enzyme activator activity / cellular response to transforming growth factor beta stimulus / heat shock protein binding / intrinsic apoptotic signaling pathway / cellular response to dexamethasone stimulus / response to nutrient / kidney development / neural tube closure / response to ischemia / positive regulation of apoptotic signaling pathway / NOD1/2 Signaling Pathway / protein processing / ADP binding / SH3 domain binding / activation of cysteine-type endopeptidase activity involved in apoptotic process / intrinsic apoptotic signaling pathway in response to DNA damage / cellular response to UV / positive regulation of neuron apoptotic process / response to estradiol / nervous system development / peptidase activity / regulation of apoptotic process / secretory granule lumen / neuron apoptotic process / ficolin-1-rich granule lumen / response to lipopolysaccharide / cell differentiation / response to hypoxia / positive regulation of apoptotic process / cysteine-type endopeptidase activity / nucleotide binding / apoptotic process / DNA damage response / Neutrophil degranulation / protein kinase binding / protein-containing complex / mitochondrion / extracellular exosome / extracellular region / ATP binding / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MIR / Resolution: 2.5 Å MIR / Resolution: 2.5 Å | ||||||
 Authors Authors | Qin, H. / Srinivasula, S. / Wu, G. / Fernandes-Alnemri, T. / Alnemri, E. / Shi, Y. | ||||||
 Citation Citation |  Journal: Nature / Year: 1999 Journal: Nature / Year: 1999Title: Structural basis of procaspase-9 recruitment by the apoptotic protease-activating factor 1. Authors: Qin, H. / Srinivasula, S.M. / Wu, G. / Fernandes-Alnemri, T. / Alnemri, E.S. / Shi, Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3ygs.cif.gz 3ygs.cif.gz | 53.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3ygs.ent.gz pdb3ygs.ent.gz | 39.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3ygs.json.gz 3ygs.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  3ygs_validation.pdf.gz 3ygs_validation.pdf.gz | 369.2 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  3ygs_full_validation.pdf.gz 3ygs_full_validation.pdf.gz | 375.2 KB | Display | |
| Data in XML |  3ygs_validation.xml.gz 3ygs_validation.xml.gz | 6 KB | Display | |
| Data in CIF |  3ygs_validation.cif.gz 3ygs_validation.cif.gz | 9.4 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/yg/3ygs https://data.pdbj.org/pub/pdb/validation_reports/yg/3ygs ftp://data.pdbj.org/pub/pdb/validation_reports/yg/3ygs ftp://data.pdbj.org/pub/pdb/validation_reports/yg/3ygs | HTTPS FTP |
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 10925.555 Da / Num. of mol.: 1 / Fragment: CASPASE RECRUITMENT DOMAIN (CARD) Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Cellular location: CYTOPLASM / Plasmid: PGEX-2T / Species (production host): Escherichia coli / Production host: Homo sapiens (human) / Cellular location: CYTOPLASM / Plasmid: PGEX-2T / Species (production host): Escherichia coli / Production host:  |
|---|---|
| #2: Protein | Mass: 11312.906 Da / Num. of mol.: 1 / Fragment: PRODOMAIN Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Cellular location: CYTOPLASM / Plasmid: PGEX-2T / Species (production host): Escherichia coli / Production host: Homo sapiens (human) / Cellular location: CYTOPLASM / Plasmid: PGEX-2T / Species (production host): Escherichia coli / Production host:  References: UniProt: P55211, Hydrolases; Acting on peptide bonds (peptidases); Cysteine endopeptidases |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.8 Å3/Da / Density % sol: 65 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 7.5 / Details: pH 7.5 | ||||||||||||||||||||
| Crystal | *PLUS | ||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / Method: vapor diffusion, hanging drop | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 |
| Detector | Details: MIRRORS |
| Radiation | Monochromator: NI FILTER / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.5→99 Å / Num. obs: 134080 / % possible obs: 98.6 % / Redundancy: 11 % / Rmerge(I) obs: 0.056 |
| Reflection shell | Resolution: 2.5→2.59 Å / Rmerge(I) obs: 0.177 / % possible all: 93.2 |
| Reflection | *PLUS Num. obs: 12452 / Num. measured all: 134080 |
| Reflection shell | *PLUS % possible obs: 93.2 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MIR / Resolution: 2.5→10 Å / σ(F): 0 MIR / Resolution: 2.5→10 Å / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.5→10 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.5→2.61 Å / Total num. of bins used: 10
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.8 / Classification: refinement X-PLOR / Version: 3.8 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.5 Å / Lowest resolution: 10 Å / σ(F): 0 / % reflection Rfree: 5 % / Rfactor obs: 0.224 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Highest resolution: 2.5 Å / Rfactor Rfree: 0.46 / % reflection Rfree: 5 % / Rfactor Rwork: 0.35 |
 Movie
Movie Controller
Controller













 PDBj
PDBj












