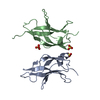
Entry Database : PDB / ID : 3jv4Title Crystal structure of the dimerization domains p50 and RelB Nuclear factor NF-kappa-B p105 subunit Transcription factor RelB Keywords / / / / / / / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Mus musculus (house mouse)Method / / Resolution : 3.15 Å Authors Vu, D. / Huang, D.B. / Ghosh, G. Journal : J.Mol.Biol. / Year : 2013Title : A structural basis for selective dimerization by NF-kappa B RelB.Authors : Vu, D. / Huang, D.B. / Vemu, A. / Ghosh, G. History Deposition Sep 15, 2009 Deposition site / Processing site Revision 1.0 Nov 24, 2010 Provider / Type Revision 1.1 Jul 13, 2011 Group Revision 1.2 Mar 27, 2013 Group Revision 1.3 Sep 4, 2013 Group Revision 1.4 Feb 21, 2024 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / struct_ncs_dom_lim Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _struct_ncs_dom_lim.beg_auth_comp_id / _struct_ncs_dom_lim.beg_label_comp_id / _struct_ncs_dom_lim.beg_label_seq_id / _struct_ncs_dom_lim.end_auth_comp_id / _struct_ncs_dom_lim.end_label_comp_id / _struct_ncs_dom_lim.end_label_seq_id
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 3.15 Å
MOLECULAR REPLACEMENT / Resolution: 3.15 Å  Authors
Authors Citation
Citation Journal: J.Mol.Biol. / Year: 2013
Journal: J.Mol.Biol. / Year: 2013 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 3jv4.cif.gz
3jv4.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb3jv4.ent.gz
pdb3jv4.ent.gz PDB format
PDB format 3jv4.json.gz
3jv4.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/jv/3jv4
https://data.pdbj.org/pub/pdb/validation_reports/jv/3jv4 ftp://data.pdbj.org/pub/pdb/validation_reports/jv/3jv4
ftp://data.pdbj.org/pub/pdb/validation_reports/jv/3jv4 Links
Links Assembly
Assembly



 Components
Components



 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 Å
ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 Å Processing
Processing MOLECULAR REPLACEMENT / Resolution: 3.15→29.76 Å / Rfactor Rfree error: 0.011 / Occupancy max: 1 / Occupancy min: 1 / FOM work R set: 0.836 / Data cutoff high absF: 32597 / Data cutoff low absF: 0 / Isotropic thermal model: GROUP / Cross valid method: THROUGHOUT / σ(F): 2 / Stereochemistry target values: Engh & Huber
MOLECULAR REPLACEMENT / Resolution: 3.15→29.76 Å / Rfactor Rfree error: 0.011 / Occupancy max: 1 / Occupancy min: 1 / FOM work R set: 0.836 / Data cutoff high absF: 32597 / Data cutoff low absF: 0 / Isotropic thermal model: GROUP / Cross valid method: THROUGHOUT / σ(F): 2 / Stereochemistry target values: Engh & Huber Movie
Movie Controller
Controller













 PDBj
PDBj





















