[English] 日本語
 Yorodumi
Yorodumi- PDB-3dpo: Crystal structure of the substrate binding domain of E. coli DnaK... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3dpo | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of the substrate binding domain of E. coli DnaK in complex with a short pyrrhocoricin-derived inhibitor peptide | ||||||
 Components Components |
| ||||||
 Keywords Keywords | Chaperone / Peptide Binding Protein / molecular chaperone / dnaK / Hsp70 / substrate-binding domain / pyrrhocoricin inhibitor / ATP-binding / Cytoplasm / DNA replication / Membrane / Nucleotide-binding / Phosphoprotein / Stress response | ||||||
| Function / homology |  Function and homology information Function and homology informationstress response to copper ion / sigma factor antagonist activity / protein unfolding / cellular response to unfolded protein / heat shock protein binding / protein folding chaperone / inclusion body / ATP-dependent protein folding chaperone / ADP binding / : ...stress response to copper ion / sigma factor antagonist activity / protein unfolding / cellular response to unfolded protein / heat shock protein binding / protein folding chaperone / inclusion body / ATP-dependent protein folding chaperone / ADP binding / : / response to heat / protein refolding / protein-folding chaperone binding / protein folding / protein-containing complex assembly / DNA replication / defense response to bacterium / innate immune response / ATP hydrolysis activity / protein-containing complex / extracellular region / zinc ion binding / ATP binding / membrane / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 2.1 Å SYNCHROTRON / Resolution: 2.1 Å | ||||||
 Authors Authors | Roujeinikova, A. | ||||||
 Citation Citation |  Journal: J.Bacteriol. / Year: 2009 Journal: J.Bacteriol. / Year: 2009Title: Allosteric coupling between the lid and interdomain linker in DnaK revealed by inhibitor binding studies Authors: Liebscher, M. / Roujeinikova, A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3dpo.cif.gz 3dpo.cif.gz | 112.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3dpo.ent.gz pdb3dpo.ent.gz | 85.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3dpo.json.gz 3dpo.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dp/3dpo https://data.pdbj.org/pub/pdb/validation_reports/dp/3dpo ftp://data.pdbj.org/pub/pdb/validation_reports/dp/3dpo ftp://data.pdbj.org/pub/pdb/validation_reports/dp/3dpo | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3dppC 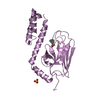 3dpqC  1dkxS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 23820.777 Da / Num. of mol.: 2 Fragment: Substrate binding domain (UNP residues 389 to 607) Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Protein/peptide | Mass: 1356.652 Da / Num. of mol.: 2 / Source method: obtained synthetically / Details: The peptide was pyrrhocoricin-derived. / References: UniProt: P37362*PLUS #3: Chemical | ChemComp-SO4 / #4: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.77 Å3/Da / Density % sol: 55.57 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 5 Details: 2.4 M ammonium sulfate, 100 mM citric acid, pH 5.0, VAPOR DIFFUSION, SITTING DROP, temperature 293K |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID23-1 / Wavelength: 1 Å / Beamline: ID23-1 / Wavelength: 1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Date: Nov 3, 2007 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.1→69.843 Å / Num. obs: 32942 / % possible obs: 98.3 % / Redundancy: 3.4 % / Rmerge(I) obs: 0.09 / Rsym value: 0.09 / Net I/σ(I): 7.1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell |
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Starting model: PDB entry 1dkx Resolution: 2.1→15 Å / Cor.coef. Fo:Fc: 0.944 / Cor.coef. Fo:Fc free: 0.898 / Occupancy max: 1 / Occupancy min: 0.5 / SU B: 4.232 / SU ML: 0.115 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.199 / ESU R Free: 0.187 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES: REFINED INDIVIDUALLY
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 61.51 Å2 / Biso mean: 22.094 Å2 / Biso min: 2 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.1→15 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.1→2.153 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller












 PDBj
PDBj



