+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3dox | ||||||
|---|---|---|---|---|---|---|---|
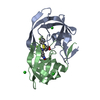
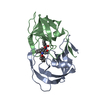
| Title | X-ray structure of HIV-1 protease in situ product complex | ||||||
 Components Components |
| ||||||
 Keywords Keywords | HYDROLASE / HIV-1 protease / transition state / reaction intermediate / catalysis / inhibitor / X-ray Crystallography / AIDS / Aspartyl protease / Capsid maturation / Capsid protein / Cytoplasm / DNA integration / DNA recombination / DNA-directed DNA polymerase / Endonuclease / Host-virus interaction / Lipoprotein / Magnesium / Metal-binding / Multifunctional enzyme / Myristate / Nuclease / Nucleotidyltransferase / Nucleus / Phosphoprotein / Protease / Ribosomal frameshifting / RNA-binding / RNA-directed DNA polymerase / Transferase / Viral nucleoprotein / Virion / Zinc / Zinc-finger | ||||||
| Function / homology |  Function and homology information Function and homology informationintegrase activity / Integration of viral DNA into host genomic DNA / Autointegration results in viral DNA circles / Minus-strand DNA synthesis / Plus-strand DNA synthesis / Uncoating of the HIV Virion / 2-LTR circle formation / Vpr-mediated nuclear import of PICs / Early Phase of HIV Life Cycle / Integration of provirus ...integrase activity / Integration of viral DNA into host genomic DNA / Autointegration results in viral DNA circles / Minus-strand DNA synthesis / Plus-strand DNA synthesis / Uncoating of the HIV Virion / 2-LTR circle formation / Vpr-mediated nuclear import of PICs / Early Phase of HIV Life Cycle / Integration of provirus / APOBEC3G mediated resistance to HIV-1 infection / Binding and entry of HIV virion / viral life cycle / HIV-1 retropepsin / symbiont-mediated activation of host apoptosis / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / Assembly Of The HIV Virion / Budding and maturation of HIV virion / protein processing / viral genome integration into host DNA / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding / viral penetration into host nucleus / host multivesicular body / RNA-directed DNA polymerase activity / RNA-DNA hybrid ribonuclease activity / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / peptidase activity / host cell / viral nucleocapsid / DNA recombination / DNA-directed DNA polymerase / aspartic-type endopeptidase activity / Hydrolases; Acting on ester bonds / DNA-directed DNA polymerase activity / symbiont-mediated suppression of host gene expression / viral translational frameshifting / symbiont entry into host cell / lipid binding / host cell plasma membrane / host cell nucleus / virion membrane / structural molecule activity / DNA binding / zinc ion binding / identical protein binding Similarity search - Function | ||||||
| Biological species |   Human immunodeficiency virus type 1 Human immunodeficiency virus type 1 | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 2 Å SYNCHROTRON / Resolution: 2 Å | ||||||
 Authors Authors | Hosur, M.V. / Ferrer, J.-L. / Das, A. / Prashar, V. / Bihani, S. | ||||||
 Citation Citation |  Journal: Proteins / Year: 2009 Journal: Proteins / Year: 2009Title: X-ray structure of HIV-1 protease in situ product complex Authors: Bihani, S. / Das, A. / Prashar, V. / Ferrer, J.-L. / Hosur, M.V. #1:  Journal: Proteins / Year: 2001 Journal: Proteins / Year: 2001Title: 1.9 A x-ray study shows closed flap conformation in crystals of tethered HIV-1 PR Authors: Pillai, B. / Kannan, K.K. / Hosur, M.V. #2: Journal: Biochem.J. / Year: 2005 Title: Observation of a tetrahedral reaction intermediate in the HIV-1 protease-substrate complex Authors: Kumar, M. / Prashar, V. / Mahale, S. / Hosur, M.V. #3: Journal: Acta Crystallogr.,Sect.D / Year: 2004 Title: Rapid screening for HIV-1 protease inhibitor leads through X-ray diffraction Authors: Pillai, B. / Kannan, K.K. / Bhat, S.V. / Hosur, M.V. #4:  Journal: Proc.Natl.Acad.Sci.Usa / Year: 2006 Journal: Proc.Natl.Acad.Sci.Usa / Year: 2006Title: Crystal structure of HIV-1 protease in situ product complex and observation of a low-barrier hydrogen bond between catalytic aspartates Authors: Das, A. / Prashar, V. / Mahale, S. / Serre, L. / Ferrer, J.-L. / Hosur, M.V. | ||||||
| History |
| ||||||
| Remark 600 | Some waters and the short peptide partially occupy the electron density. In some unit cells the ...Some waters and the short peptide partially occupy the electron density. In some unit cells the water cluster is there while in others the ligand is present. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3dox.cif.gz 3dox.cif.gz | 57.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3dox.ent.gz pdb3dox.ent.gz | 41.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3dox.json.gz 3dox.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/do/3dox https://data.pdbj.org/pub/pdb/validation_reports/do/3dox ftp://data.pdbj.org/pub/pdb/validation_reports/do/3dox ftp://data.pdbj.org/pub/pdb/validation_reports/do/3dox | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 21902.760 Da / Num. of mol.: 1 / Mutation: C95M, C1095A Source method: isolated from a genetically manipulated source Details: TETHERED DIMER LINKED BY GGSSG Source: (gene. exp.)  Human immunodeficiency virus type 1 (isolate HXB2 group M subtype B) Human immunodeficiency virus type 1 (isolate HXB2 group M subtype B)Gene: gag-pol / Production host:  |
|---|---|
| #2: Protein/peptide | Mass: 510.498 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: The peptide was chemically synthesized. |
| #3: Protein/peptide | Mass: 327.419 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: The peptide was chemically synthesized. |
| #4: Water | ChemComp-HOH / |
| Sequence details | THIS CONSTRUCT INCLUDES A LINKER THAT CONSIST OF 996TH GLY, 997TH GLY, 998TH SER, 999TH SER AND 1000TH GLY. |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.03 Å3/Da / Density % sol: 39.27 % |
|---|---|
| Crystal grow | Temperature: 300 K / pH: 6.2 Details: 1-5% saturated ammonium sulfate, pH 6.2, Soaking, temperature 300.0K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SLS SLS  / Beamline: X06SA / Wavelength: 0.97644 Å / Beamline: X06SA / Wavelength: 0.97644 Å |
| Detector | Type: MARMOSAIC 225 mm CCD / Detector: CCD / Date: Dec 6, 2006 |
| Radiation | Monochromator: Double Crystal Monochromator / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.97644 Å / Relative weight: 1 |
| Reflection | Resolution: 2→50 Å / Num. all: 12272 / Num. obs: 12068 / % possible obs: 98.3 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Rmerge(I) obs: 0.08 / Net I/σ(I): 18.03 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2→45.11 Å / Occupancy max: 1 / Occupancy min: 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Bsol: 77.83 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 34.13 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→45.11 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller























 PDBj
PDBj








