| Entry | Database: PDB / ID: 2vtg
|
|---|
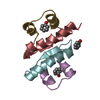
| Title | Crystal Structure of Human Iba2, trigonal crystal form |
|---|
 Components Components | IONIZED CALCIUM-BINDING ADAPTER MOLECULE 2 |
|---|
 Keywords Keywords | METAL BINDING PROTEIN / EF-HAND / CALCIUM BINDING / ACTIN CROSSLINKING / IONIZED CALCIUM BINDING ADAPTER MOLECULE 2 / METAL-BINDING PROTEIN |
|---|
| Function / homology |  Function and homology information Function and homology information
ruffle assembly / actin filament bundle assembly / ruffle membrane / actin filament binding / actin cytoskeleton / focal adhesion / calcium ion binding / extracellular exosomeSimilarity search - Function Allograft inflammatory factor 1 / : / Allograft inflammatory factor 1 / EF-hand calcium-binding domain profile. / EF-hand domain / EF-hand domain pairSimilarity search - Domain/homology |
|---|
| Biological species |  HOMO SAPIENS (human) HOMO SAPIENS (human) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.45 Å MOLECULAR REPLACEMENT / Resolution: 2.45 Å |
|---|
 Authors Authors | Schulze, J.O. / Quedenau, C. / Roske, Y. / Turnbull, A. / Mueller, U. / Heinemann, U. / Buessow, K. |
|---|
 Citation Citation |  Journal: FEBS J. / Year: 2008 Journal: FEBS J. / Year: 2008
Title: Structural and Functional Characterization of Human Iba Proteins.
Authors: Schulze, J.O. / Quedenau, C. / Roske, Y. / Adam, T. / Schuler, H. / Behlke, J. / Turnbull, A.P. / Sievert, V. / Scheich, C. / Mueller, U. / Heinemann, U. / Bussow, K. |
|---|
| History | | Deposition | May 15, 2008 | Deposition site: PDBE / Processing site: PDBE |
|---|
| Revision 1.0 | Jul 14, 2009 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Jul 13, 2011 | Group: Advisory / Version format compliance |
|---|
| Revision 1.2 | Dec 13, 2023 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Other / Refinement description
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / database_2 / pdbx_database_status / pdbx_initial_refinement_model / pdbx_struct_conn_angle / struct_conn / struct_site
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession ..._database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_sf / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr1_symmetry / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.ptnr3_symmetry / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr1_symmetry / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_conn.ptnr2_symmetry / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id |
|---|
| Revision 1.3 | Oct 23, 2024 | Group: Structure summary / Category: pdbx_entry_details / pdbx_modification_feature |
|---|
|
|---|
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information HOMO SAPIENS (human)
HOMO SAPIENS (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.45 Å
MOLECULAR REPLACEMENT / Resolution: 2.45 Å  Authors
Authors Citation
Citation Journal: FEBS J. / Year: 2008
Journal: FEBS J. / Year: 2008 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2vtg.cif.gz
2vtg.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2vtg.ent.gz
pdb2vtg.ent.gz PDB format
PDB format 2vtg.json.gz
2vtg.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/vt/2vtg
https://data.pdbj.org/pub/pdb/validation_reports/vt/2vtg ftp://data.pdbj.org/pub/pdb/validation_reports/vt/2vtg
ftp://data.pdbj.org/pub/pdb/validation_reports/vt/2vtg

 Links
Links Assembly
Assembly
 Components
Components HOMO SAPIENS (human) / Production host:
HOMO SAPIENS (human) / Production host: 
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  BESSY
BESSY  / Beamline: 14.1 / Wavelength: 0.9184
/ Beamline: 14.1 / Wavelength: 0.9184  Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller












 PDBj
PDBj



