| Entry | Database: PDB / ID: 2v94
|
|---|
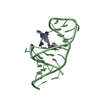
| Title | Crystal structure of P. abyssi RPS24 |
|---|
 Components Components | 30S RIBOSOMAL PROTEIN S24E |
|---|
 Keywords Keywords | RIBOSOMAL PROTEIN / RIBONUCLEOPROTEIN |
|---|
| Function / homology |  Function and homology information Function and homology information
RRM (RNA recognition motif) domain / Ribosomal S24e conserved site / Ribosomal protein S24e signature. / Ribosomal protein S24e / Ribosomal protein S24e / Ribosomal protein L23/L15e core domain superfamily / Nucleotide-binding alpha-beta plait domain superfamily / Alpha-Beta Plaits / 2-Layer Sandwich / Alpha BetaSimilarity search - Domain/homology |
|---|
| Biological species |   PYROCOCCUS ABYSSI (archaea) PYROCOCCUS ABYSSI (archaea) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 1.9 Å SAD / Resolution: 1.9 Å |
|---|
 Authors Authors | Legrand, P. / Pinaud, N. / Gleizes, P.E. / Fribourg, S. |
|---|
 Citation Citation |  Journal: Hum.Mol.Genet. / Year: 2008 Journal: Hum.Mol.Genet. / Year: 2008
Title: Mutation of Ribosomal Protein Rps24 in Diamond- Blackfan Anemia Results in a Ribosome Biogenesis Disorder.
Authors: Choesmel, V. / Fribourg, S. / Aguissa-Toure, A.H. / Pinaud, N. / Legrand, P. / Gazda, H.T. / Gleizes, P.E. |
|---|
| History | | Deposition | Aug 21, 2007 | Deposition site: PDBE / Processing site: PDBE |
|---|
| Revision 1.0 | Apr 29, 2008 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Jul 13, 2011 | Group: Advisory / Refinement description / Version format compliance |
|---|
| Revision 1.2 | Nov 6, 2024 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Other / Refinement description / Structure summary
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / database_2 / pdbx_database_status / pdbx_entry_details / pdbx_modification_feature / struct_conn / struct_ncs_dom_lim
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession ..._database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_sf / _struct_conn.pdbx_leaving_atom_flag / _struct_ncs_dom_lim.beg_auth_comp_id / _struct_ncs_dom_lim.beg_label_asym_id / _struct_ncs_dom_lim.beg_label_comp_id / _struct_ncs_dom_lim.beg_label_seq_id / _struct_ncs_dom_lim.end_auth_comp_id / _struct_ncs_dom_lim.end_label_asym_id / _struct_ncs_dom_lim.end_label_comp_id / _struct_ncs_dom_lim.end_label_seq_id |
|---|
|
|---|
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 PYROCOCCUS ABYSSI (archaea)
PYROCOCCUS ABYSSI (archaea) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  SAD / Resolution: 1.9 Å
SAD / Resolution: 1.9 Å  Authors
Authors Citation
Citation Journal: Hum.Mol.Genet. / Year: 2008
Journal: Hum.Mol.Genet. / Year: 2008 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2v94.cif.gz
2v94.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2v94.ent.gz
pdb2v94.ent.gz PDB format
PDB format 2v94.json.gz
2v94.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/v9/2v94
https://data.pdbj.org/pub/pdb/validation_reports/v9/2v94 ftp://data.pdbj.org/pub/pdb/validation_reports/v9/2v94
ftp://data.pdbj.org/pub/pdb/validation_reports/v9/2v94 Links
Links Assembly
Assembly


 Movie
Movie Controller
Controller












 PDBj
PDBj