[English] 日本語
 Yorodumi
Yorodumi- PDB-2v86: Crystal structure of RAG2-PHD finger in complex with H3R2me2aK4me... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2v86 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
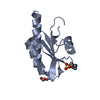
| Title | Crystal structure of RAG2-PHD finger in complex with H3R2me2aK4me3 peptide | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords | PROTEIN BINDING / V(D)J RECOMBINATION / COVALENT MODIFICATIONS / RAG2 / HISTONE / NUCLEUS / NUCLEASE / HYDROLASE / PHD FINGER / DNA-BINDING / RECOMBINASE / ENDONUCLEASE / TRIMETHYL LYSINE / DIMETHYL ARGININE / DNA RECOMBINATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationChromatin modifying enzymes / Interleukin-7 signaling / Negative Regulation of CDH1 Gene Transcription / Regulation of endogenous retroelements by KRAB-ZFP proteins / Condensation of Prophase Chromosomes / HDMs demethylate histones / HDACs deacetylate histones / PRC2 methylates histones and DNA / mature B cell differentiation involved in immune response / HATs acetylate histones ...Chromatin modifying enzymes / Interleukin-7 signaling / Negative Regulation of CDH1 Gene Transcription / Regulation of endogenous retroelements by KRAB-ZFP proteins / Condensation of Prophase Chromosomes / HDMs demethylate histones / HDACs deacetylate histones / PRC2 methylates histones and DNA / mature B cell differentiation involved in immune response / HATs acetylate histones / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / PKMTs methylate histone lysines / DNA recombinase complex / B cell homeostatic proliferation / negative regulation of T cell differentiation in thymus / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / RMTs methylate histone arginines / pre-B cell allelic exclusion / Factors involved in megakaryocyte development and platelet production / Estrogen-dependent gene expression / positive regulation of organ growth / V(D)J recombination / phosphatidylinositol-3,4-bisphosphate binding / histone H3K4me3 reader activity / phosphatidylinositol-3,5-bisphosphate binding / T cell lineage commitment / B cell lineage commitment / phosphatidylinositol-3,4,5-trisphosphate binding / T cell differentiation / phosphatidylinositol-4,5-bisphosphate binding / phosphatidylinositol binding / epigenetic regulation of gene expression / B cell differentiation / structural constituent of chromatin / T cell differentiation in thymus / nucleosome / nucleosome assembly / chromatin organization / DNA recombination / sequence-specific DNA binding / gene expression / defense response to bacterium / protein heterodimerization activity / chromatin binding / chromatin / negative regulation of transcription by RNA polymerase II / DNA binding / zinc ion binding / nucleoplasm / nucleus Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.05 Å MOLECULAR REPLACEMENT / Resolution: 2.05 Å | |||||||||
 Authors Authors | Ramon-Maiques, S. / Yang, W. | |||||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 2007 Journal: Proc.Natl.Acad.Sci.USA / Year: 2007Title: The Plant Homeodomain Finger of Rag2 Recognizes Histone H3 Methylated at Both Lysine-4 and Arginine-2. Authors: Ramon-Maiques, S. / Kuo, A.J. / Carney, D. / Matthews, A.G.W. / Oettinger, M.A. / Gozani, O. / Yang, W. #1:  Journal: Nature / Year: 2007 Journal: Nature / Year: 2007Title: Rag2 Phd Finger Couples Histone H3 Lysine 4 Trimethylation with V(D)J Recombination. Authors: Matthews, A.G.W. / Kuo, A.J. / Ramon-Maiques, S. / Han, S. / Champagne, K.S. / Ivanov, D. / Gallardo, M. / Carney, D. / Cheung, P. / Ciccone, D.N. / Walter, K.L. / Utz, P.J. / Shi, Y. / ...Authors: Matthews, A.G.W. / Kuo, A.J. / Ramon-Maiques, S. / Han, S. / Champagne, K.S. / Ivanov, D. / Gallardo, M. / Carney, D. / Cheung, P. / Ciccone, D.N. / Walter, K.L. / Utz, P.J. / Shi, Y. / Kutateladze, T.G. / Yang, W. / Gozani, O. / Oettinger, M.A. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2v86.cif.gz 2v86.cif.gz | 60.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2v86.ent.gz pdb2v86.ent.gz | 43.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2v86.json.gz 2v86.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/v8/2v86 https://data.pdbj.org/pub/pdb/validation_reports/v8/2v86 ftp://data.pdbj.org/pub/pdb/validation_reports/v8/2v86 ftp://data.pdbj.org/pub/pdb/validation_reports/v8/2v86 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2v83SC  2v85C  2v87C  2v88C S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 9360.498 Da / Num. of mol.: 2 / Fragment: RESIDUES 414-487 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Protein/peptide | Mass: 1074.277 Da / Num. of mol.: 2 / Fragment: H3 (1-21), BIOTINILATED AT C-TERMINUS / Source method: obtained synthetically Details: R2 ASYMMETRICALLY DIMETHYLATED AND K4 TRIMETHYLATED Source: (synth.)  #3: Chemical | ChemComp-ZN / #4: Water | ChemComp-HOH / | Sequence details | N-TERMINAL SEGMENT GPLGSPEFG IS CARRIED OVER FROM THE EXPRESSION | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.23 Å3/Da / Density % sol: 44.48 % / Description: NONE |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop Details: VAPOR DIFFUSION. HANGING DROP. 28% PEG 20,000, 0.1M MES PH 6.5, 3% ISOPROPANOL TEMPERATURE 293K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 |
| Detector | Type: RIGAKU-MSC / Detector: IMAGE PLATE / Date: Mar 4, 2006 / Details: MIRRORS |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.05→30 Å / Num. obs: 11827 / % possible obs: 99.8 % / Observed criterion σ(I): 1 / Redundancy: 5 % / Biso Wilson estimate: 18.5 Å2 / Rmerge(I) obs: 0.18 / Net I/σ(I): 9.9 |
| Reflection shell | Resolution: 2.05→2.12 Å / Redundancy: 5 % / Rmerge(I) obs: 0.52 / Mean I/σ(I) obs: 3 / % possible all: 99.9 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2V83 Resolution: 2.05→30 Å / Rfactor Rfree error: 0.00087 / Data cutoff high absF: 10000 / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: MAXIMUM LIKELIHOOD
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 47.2446 Å2 / ksol: 0.360332 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 18.496 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.21 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.05→30 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.05→2.12 Å / Rfactor Rfree error: 0.04 / Total num. of bins used: 11
|
 Movie
Movie Controller
Controller













 PDBj
PDBj









