+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2qlg | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
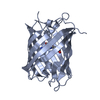
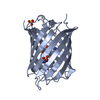
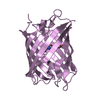
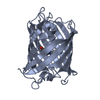
| Title | mPlum | |||||||||
 Components Components | (Fluorescent protein plum) x 2 | |||||||||
 Keywords Keywords | FLUORESCENT PROTEIN / far-red fluorescent protein / acylimine | |||||||||
| Function / homology |  Function and homology information Function and homology information | |||||||||
| Biological species |  Discosoma sp. LW-2004 (sea anemone) Discosoma sp. LW-2004 (sea anemone) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å | |||||||||
 Authors Authors | Shu, X. / Remington, S.J. | |||||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: Structural Studies of Far-Red Emission in mPlum, a Monomeric Red Fluorescent Protein Authors: Shu, X. / Wang, L. / Colip, L. / Kallio, K. / Remington, S.J. | |||||||||
| History |
| |||||||||
| Remark 400 | COMPOUND RELATED TO PROTEIN THE TWO CHAINS IN ASSYMETRIC UNIT HAD THE SAME SEQUENCE EXCEPT AT THE ...COMPOUND RELATED TO PROTEIN THE TWO CHAINS IN ASSYMETRIC UNIT HAD THE SAME SEQUENCE EXCEPT AT THE CHROMPHORE. THE CH6 OF CHAIN A AND NRQ OF CHAIN B BOTH COME FROM THE SAME PARENT RESIDUES MET, TYR AND GLY. AUTHOR STATED THAT THE CA1-N BOND OF CH6 IS SINGLE BOND BUT THE CA1-N OF NRQ IS DOUBLE BOND THAT EXTENDS THE CHROMOPHORE CONJUGATION SYSTEM. THIS IS THE ONLY DIFFERENCE BETWEEN CR2 AND CR3. AUTHOR ALSO STATED THAT CR2 IS A RED FLUORESCENT PROTEIN CHROMOPHORE, WHILE CR3 IS A GREEN FLUORESCENT PROTEIN CHROMOPHORE, DUE TO DIFFERENT CONJUGATION LENGTH. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2qlg.cif.gz 2qlg.cif.gz | 109.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2qlg.ent.gz pdb2qlg.ent.gz | 82.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2qlg.json.gz 2qlg.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ql/2qlg https://data.pdbj.org/pub/pdb/validation_reports/ql/2qlg ftp://data.pdbj.org/pub/pdb/validation_reports/ql/2qlg ftp://data.pdbj.org/pub/pdb/validation_reports/ql/2qlg | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2qlhC  2qliC  1g7kS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 25606.928 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Discosoma sp. LW-2004 (sea anemone) / Plasmid: pBAD / Production host: Discosoma sp. LW-2004 (sea anemone) / Plasmid: pBAD / Production host:  |
|---|---|
| #2: Protein | Mass: 25604.912 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Discosoma sp. LW-2004 (sea anemone) / Production host: Discosoma sp. LW-2004 (sea anemone) / Production host:  |
| #3: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.26 Å3/Da / Density % sol: 45.52 % |
|---|---|
| Crystal grow | pH: 8.5 / Details: 200mM NaCl, 100mM Tris pH8.5, 30% PEG 3400 |
-Data collection
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU / Wavelength: 1.54 Å ROTATING ANODE / Type: RIGAKU / Wavelength: 1.54 Å |
|---|---|
| Detector | Date: Sep 25, 2005 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.54 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→10 Å / Num. obs: 43227 |
- Processing
Processing
| Software |
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB entry 1G7K Resolution: 1.8→10 Å
| |||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→10 Å
|
 Movie
Movie Controller
Controller




















 PDBj
PDBj


