[English] 日本語
 Yorodumi
Yorodumi- PDB-2q71: Uroporphyrinogen Decarboxylase G168R single mutant enzyme in comp... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2q71 | ||||||
|---|---|---|---|---|---|---|---|
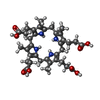
| Title | Uroporphyrinogen Decarboxylase G168R single mutant enzyme in complex with coproporphyrinogen-III | ||||||
 Components Components | Uroporphyrinogen decarboxylase | ||||||
 Keywords Keywords | LYASE / Uroporphyrinogen Decarboxylase enzyme UROD G168R coproporphyrinogen-III product complex | ||||||
| Function / homology |  Function and homology information Function and homology informationporphyrin-containing compound catabolic process / uroporphyrinogen decarboxylase / uroporphyrinogen decarboxylase activity / porphyrin-containing compound metabolic process / heme A biosynthetic process / heme B biosynthetic process / : / Heme biosynthesis / heme biosynthetic process / nucleoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.9 Å MOLECULAR REPLACEMENT / Resolution: 1.9 Å | ||||||
 Authors Authors | Phillips, J.D. / Whitby, F.G. / Stadtmueller, B.M. / Edwards, C.Q. / Hill, C.P. / Kushner, J.P. | ||||||
 Citation Citation |  Journal: Transl.Res. / Year: 2007 Journal: Transl.Res. / Year: 2007Title: Two Novel Uropophyrinogen Decarboxylase (URO-D) Mutations Causing Hepatoerythropoietic Porphyria (HEP) Authors: Phillips, J.D. / Whitby, F.G. / Stadtmueller, B.M. / Edwards, C.Q. / Hill, C.P. / Kushner, J.P. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2q71.cif.gz 2q71.cif.gz | 98.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2q71.ent.gz pdb2q71.ent.gz | 73.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2q71.json.gz 2q71.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/q7/2q71 https://data.pdbj.org/pub/pdb/validation_reports/q7/2q71 ftp://data.pdbj.org/pub/pdb/validation_reports/q7/2q71 ftp://data.pdbj.org/pub/pdb/validation_reports/q7/2q71 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2q6zC  1ry3S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | The crystal contains one monomer per asymmetric unit. The functional enzyme is a dimer of identical subunits. A crystallographic 2-fold generates teh biologically functional dimeric enzyme. The two identical active sites have been shown to act independently. One reaction product molecule (copropophyrinogen-III) is in the active site. |
- Components
Components
| #1: Protein | Mass: 39862.770 Da / Num. of mol.: 1 / Mutation: G168R Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: UROD / Plasmid: pET-16b / Production host: Homo sapiens (human) / Gene: UROD / Plasmid: pET-16b / Production host:  |
|---|---|
| #2: Chemical | ChemComp-CP3 / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.76 Å3/Da / Density % sol: 55.45 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, sitting drop / pH: 7 Details: Protien at 6.5 mg/ml in 50mM Tris, pH 7.5, 1mM BME was mixed 5 parts to 3 parts of precipitant (1.7M citrate, pH 7.0) and equilibrated by sitting drop vapor diffusion against a well of ...Details: Protien at 6.5 mg/ml in 50mM Tris, pH 7.5, 1mM BME was mixed 5 parts to 3 parts of precipitant (1.7M citrate, pH 7.0) and equilibrated by sitting drop vapor diffusion against a well of precipitant., VAPOR DIFFUSION, SITTING DROP, temperature 298K |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: RIGAKU RAXIS IV / Detector: IMAGE PLATE / Date: Sep 19, 2003 / Details: Yale focusing mirrors | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Monochromator: Yale Focusing mirrors, 1.5418 angstrom wavelength, 67 degrees of 0.5 degree oscillations (134 images, 0-67 degrees) plus 12 degrees of 0.25 degree oscillations (48 images 65-77 degrees) Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 1.9→30 Å / Num. obs: 32683 / % possible obs: 93.5 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Rmerge(I) obs: 0.078 / Χ2: 1.004 / Net I/σ(I): 11.3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell |
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1ry3 Resolution: 1.9→30 Å / Cor.coef. Fo:Fc: 0.963 / Cor.coef. Fo:Fc free: 0.951 / SU B: 2.943 / SU ML: 0.087 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.14 / ESU R Free: 0.129 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS. THE CRYSTALS WERE ISOMORPHOUS TO 1RY3.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 27.004 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.9→30 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.9→1.951 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller












 PDBj
PDBj