[English] 日本語
 Yorodumi
Yorodumi- PDB-2mtk: NMR structure of the III-IV-V three-way junction from the VS ribo... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2mtk | ||||||
|---|---|---|---|---|---|---|---|
| Title | NMR structure of the III-IV-V three-way junction from the VS ribozyme and identification of magnesium-binding sites using paramagnetic relaxation enhancement | ||||||
 Components Components | RNA (47-MER) | ||||||
 Keywords Keywords | RNA / VS ribozyme / Three-way junction / Base triple / U-turn / Ribose zipper / GAAA tetraloop / CUUG tetraloop | ||||||
| Function / homology | RNA / RNA (> 10) Function and homology information Function and homology information | ||||||
| Biological species |  Neurospora crassa (fungus) Neurospora crassa (fungus) | ||||||
| Method | SOLUTION NMR / simulated annealing | ||||||
| Model details | lowest energy, model1 | ||||||
 Authors Authors | Bonneau, E. / Legault, P. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2014 Journal: Biochemistry / Year: 2014Title: Nuclear Magnetic Resonance Structure of the III-IV-V Three-Way Junction from the Varkud Satellite Ribozyme and Identification of Magnesium-Binding Sites Using Paramagnetic Relaxation Enhancement. Authors: Bonneau, E. / Legault, P. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2mtk.cif.gz 2mtk.cif.gz | 627.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2mtk.ent.gz pdb2mtk.ent.gz | 532.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2mtk.json.gz 2mtk.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/mt/2mtk https://data.pdbj.org/pub/pdb/validation_reports/mt/2mtk ftp://data.pdbj.org/pub/pdb/validation_reports/mt/2mtk ftp://data.pdbj.org/pub/pdb/validation_reports/mt/2mtk | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 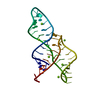
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: RNA chain | Mass: 15117.960 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  Neurospora crassa (fungus) Neurospora crassa (fungus) | ||
|---|---|---|---|
| #2: Chemical | ChemComp-MG / #3: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||
| Sample conditions |
|
-NMR measurement
| NMR spectrometer | Type: Varian Unity / Manufacturer: Varian / Model: UNITY / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: simulated annealing / Software ordinal: 1 | ||||||||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 300 / Conformers submitted total number: 21 |
 Movie
Movie Controller
Controller








 PDBj
PDBj
































 HSQC
HSQC