[English] 日本語
 Yorodumi
Yorodumi- PDB-2mrt: CONFORMATION OF CD-7 METALLOTHIONEIN-2 FROM RAT LIVER IN AQUEOUS ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2mrt | ||||||
|---|---|---|---|---|---|---|---|
| Title | CONFORMATION OF CD-7 METALLOTHIONEIN-2 FROM RAT LIVER IN AQUEOUS SOLUTION DETERMINED BY NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY | ||||||
 Components Components | CD7 METALLOTHIONEIN-2 | ||||||
 Keywords Keywords | METALLOTHIONEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationMetallothioneins bind metals / cellular response to erythropoietin / negative regulation of growth / detoxification of copper ion / intracellular zinc ion homeostasis / cellular response to interleukin-3 / cadmium ion binding / cellular response to cadmium ion / cellular response to zinc ion / : ...Metallothioneins bind metals / cellular response to erythropoietin / negative regulation of growth / detoxification of copper ion / intracellular zinc ion homeostasis / cellular response to interleukin-3 / cadmium ion binding / cellular response to cadmium ion / cellular response to zinc ion / : / astrocyte activation / cellular response to copper ion / response to bacterium / zinc ion binding / metal ion binding / nucleus / cytoplasm Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR | ||||||
 Authors Authors | Braun, W. / Schultze, P. / Woergoetter, E. / Wagner, G. / Vasak, M. / Kaegi, J.H.R. / Wuthrich, K. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1988 Journal: J.Mol.Biol. / Year: 1988Title: Conformation of [Cd7]-metallothionein-2 from rat liver in aqueous solution determined by nuclear magnetic resonance spectroscopy. Authors: Schultze, P. / Worgotter, E. / Braun, W. / Wagner, G. / Vasak, M. / Kagi, J.H. / Wuthrich, K. #1:  Journal: J.Mol.Biol. / Year: 1988 Journal: J.Mol.Biol. / Year: 1988Title: Three-Dimensional Structure of Rabbit Liver Cd-7 Metallothionein-2A in Aqueous Solution Determined by Nuclear Magnetic Resonance. Authors: Arseniev, A. / Schultze, P. / Woergoetter, E. / Braun, W. / Wagner, G. / Vasak, M. / Kaegi, J.H.R. / Wuthrich, K. #2:  Journal: Eur.J.Biochem. / Year: 1987 Journal: Eur.J.Biochem. / Year: 1987Title: Sequence-Specific 1H-NMR Assignments in Rat-Liver Metallothionein-2 Authors: Woergoetter, E. / Wagner, G. / Vasak, M. / Kaegi, J.H.R. / Wuthrich, K. #3:  Journal: J.Mol.Biol. / Year: 1987 Journal: J.Mol.Biol. / Year: 1987Title: Metal Co-Ordination in Rat Liver Metallothionein-2 Prepared with or without Reconstitution of the Metal Clusters, and Comparison with Rabbit Liver Metallothionein-2. Authors: Vasak, M. / Woergoetter, E. / Wagner, G. / Kaegi, J.H.R. / Wuthrich, K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2mrt.cif.gz 2mrt.cif.gz | 14.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2mrt.ent.gz pdb2mrt.ent.gz | 8.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2mrt.json.gz 2mrt.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/mr/2mrt https://data.pdbj.org/pub/pdb/validation_reports/mr/2mrt ftp://data.pdbj.org/pub/pdb/validation_reports/mr/2mrt ftp://data.pdbj.org/pub/pdb/validation_reports/mr/2mrt | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 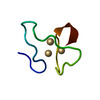
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| Atom site foot note | 1: THE THREE CADMIUM ATOMS AND THE NINE SULPHUR ATOMS TO WHICH THEY ARE BONDED FORM A CD3 S9 METAL CLUSTER. | |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 3057.594 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  |
|---|---|
| #2: Chemical |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
| Crystal grow | *PLUS Method: other / Details: NMR |
|---|
- Processing
Processing
| NMR software | Name: DISMAN / Developer: BRAUN,GO / Classification: refinement |
|---|---|
| NMR ensemble | Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller





 PDBj
PDBj

