+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2lc6 | ||||||
|---|---|---|---|---|---|---|---|
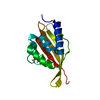
| Title | Solution structure of Par-6 Q144C/L164C | ||||||
 Components Components | Par-6 | ||||||
 Keywords Keywords | CELL ADHESION / PDZ domain / CRIB / Cdc42 | ||||||
| Function / homology |  Function and homology information Function and homology informationRAC1 GTPase cycle / RHOV GTPase cycle / terminal branching, open tracheal system / establishment or maintenance of neuroblast polarity / branching involved in open tracheal system development / zonula adherens assembly / subapical complex / TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) / establishment of neuroblast polarity / Asymmetric localization of PCP proteins ...RAC1 GTPase cycle / RHOV GTPase cycle / terminal branching, open tracheal system / establishment or maintenance of neuroblast polarity / branching involved in open tracheal system development / zonula adherens assembly / subapical complex / TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) / establishment of neuroblast polarity / Asymmetric localization of PCP proteins / RHOU GTPase cycle / muscle cell postsynaptic specialization / establishment or maintenance of polarity of embryonic epithelium / border follicle cell migration / PAR polarity complex / asymmetric neuroblast division / zonula adherens / apical cortex / establishment or maintenance of epithelial cell apical/basal polarity / morphogenesis of a polarized epithelium / positive regulation of smoothened signaling pathway / apical protein localization / positive regulation of filopodium assembly / centrosome cycle / protein kinase inhibitor activity / bicellular tight junction / positive regulation of lamellipodium assembly / synapse assembly / apical part of cell / terminal bouton / cell cortex / apical plasma membrane / plasma membrane Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR / torsion angle dynamics | ||||||
| Model details | lowest energy, model 1 | ||||||
 Authors Authors | Volkman, B.F. / Whitney, D.S. / Peterson, F.C. | ||||||
 Citation Citation |  Journal: Structure / Year: 2011 Journal: Structure / Year: 2011Title: A Conformational Switch in the CRIB-PDZ Module of Par-6. Authors: Whitney, D.S. / Peterson, F.C. / Volkman, B.F. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2lc6.cif.gz 2lc6.cif.gz | 786.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2lc6.ent.gz pdb2lc6.ent.gz | 665.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2lc6.json.gz 2lc6.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/lc/2lc6 https://data.pdbj.org/pub/pdb/validation_reports/lc/2lc6 ftp://data.pdbj.org/pub/pdb/validation_reports/lc/2lc6 ftp://data.pdbj.org/pub/pdb/validation_reports/lc/2lc6 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 13724.749 Da / Num. of mol.: 1 / Fragment: sequence database residues 130-255 / Mutation: Q144C, L164C Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 1 mM [U-100% 13C; U-100% 15N] Par-6, 20 mM sodium phosphate, 50 mM sodium chloride, 90% H2O, 10% D2O Solvent system: 90% H2O/10% D2O | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||
| Sample conditions | Ionic strength: 53 / pH: 5.5 / Pressure: AMBIENT / Temperature: 298 K |
-NMR measurement
| NMR spectrometer | Type: Bruker Avance II / Manufacturer: Bruker / Model: AVANCE II / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: torsion angle dynamics / Software ordinal: 1 Details: AUTOMATED METHODS WERE USED FOR BACKBONE CHEMICAL SHIFT ASSIGNMENT AND ITERATIVE NOE REFINEMENT. FINAL STRUCTURES WERE OBTAINED BY MOLECULAR DYNAMICS IN EXPLICIT SOLVENTSTRUCTURES ARE BASED ...Details: AUTOMATED METHODS WERE USED FOR BACKBONE CHEMICAL SHIFT ASSIGNMENT AND ITERATIVE NOE REFINEMENT. FINAL STRUCTURES WERE OBTAINED BY MOLECULAR DYNAMICS IN EXPLICIT SOLVENTSTRUCTURES ARE BASED ON A TOTAL OF 1604 NOE CONSTRAINTS ( 407 INTRA, 388 SEQUENTIAL, 251 MEDIUM, AND 558 LONG RANGE) AND 165 PHI AND PSI DIHEDRAL ANGLE CONSTRAINTS., STRUCTURES ARE BASED ON A TOTAL OF 1604 NOE CONSTRAINTS ( 407 INTRA, 388 SEQUENTIAL, 251 MEDIUM, AND 558 LONG RANGE) AND 165 PHI AND PSI DIHEDRAL ANGLE CONSTRAINTS. | ||||||||||||||||||||||||||||
| NMR constraints | NOE constraints total: 1604 / NOE intraresidue total count: 407 / NOE long range total count: 558 / NOE medium range total count: 251 / NOE sequential total count: 388 / Protein phi angle constraints total count: 82 / Protein psi angle constraints total count: 83 | ||||||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: target function / Conformers calculated total number: 100 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller













 PDBj
PDBj

 Xplor-NIH
Xplor-NIH