[English] 日本語
 Yorodumi
Yorodumi- PDB-2kfx: Structure of the N-terminal domain of human cardiac troponin C bo... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2kfx | ||||||
|---|---|---|---|---|---|---|---|
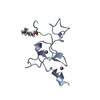
| Title | Structure of the N-terminal domain of human cardiac troponin C bound to calcium ion and to the inhibitor W7 | ||||||
 Components Components | Troponin C, slow skeletal and cardiac muscles | ||||||
 Keywords Keywords | METAL BINDING PROTEIN / calcium regulation / striated muscle / cardiac / troponin / W7 / cardiotonic drugs / Acetylation / Calcium / Cardiomyopathy / Disease mutation / Muscle protein / Polymorphism | ||||||
| Function / homology |  Function and homology information Function and homology informationdiaphragm contraction / regulation of muscle filament sliding speed / troponin T binding / cardiac Troponin complex / troponin complex / regulation of muscle contraction / transition between fast and slow fiber / Striated Muscle Contraction / muscle filament sliding / response to metal ion ...diaphragm contraction / regulation of muscle filament sliding speed / troponin T binding / cardiac Troponin complex / troponin complex / regulation of muscle contraction / transition between fast and slow fiber / Striated Muscle Contraction / muscle filament sliding / response to metal ion / cardiac muscle cell contraction / ventricular cardiac muscle tissue morphogenesis / troponin I binding / skeletal muscle contraction / cardiac muscle contraction / sarcomere / calcium-dependent protein binding / actin filament binding / calcium ion binding / protein homodimerization activity / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method | SOLUTION NMR / molecular dynamics | ||||||
 Authors Authors | Hoffman, R.M.B. / Sykes, B.D. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2009 Journal: Biochemistry / Year: 2009Title: Structure of the inhibitor W7 bound to the regulatory domain of cardiac troponin C. Authors: Hoffman, R.M. / Sykes, B.D. #1:  Journal: Biochemistry / Year: 1999 Journal: Biochemistry / Year: 1999Title: Binding of cardiac troponin-I147-163 induces a structural opening in human cardiac troponin-C. Authors: Li, M.X. / Spyracopoulos, L. / Sykes, B.D. #2:  Journal: J.Biol.Chem. / Year: 2002 Journal: J.Biol.Chem. / Year: 2002Title: Structure of the regulatory N-domain of human cardiac troponin C in complex with human cardiac troponin I147-163 and bepridil. Authors: Wang, X. / Li, M.X. / Sykes, B.D. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2kfx.cif.gz 2kfx.cif.gz | 278.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2kfx.ent.gz pdb2kfx.ent.gz | 233.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2kfx.json.gz 2kfx.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kf/2kfx https://data.pdbj.org/pub/pdb/validation_reports/kf/2kfx ftp://data.pdbj.org/pub/pdb/validation_reports/kf/2kfx ftp://data.pdbj.org/pub/pdb/validation_reports/kf/2kfx | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 10038.173 Da / Num. of mol.: 1 / Mutation: C35S, C84S Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: TNNC1, TNNC / Plasmid: pET / Production host: Homo sapiens (human) / Gene: TNNC1, TNNC / Plasmid: pET / Production host:  |
|---|---|
| #2: Chemical | ChemComp-CA / |
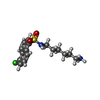
| #3: Chemical | ChemComp-WW7 / |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR Details: Structure of the N-terminal domain of human cardiac troponin C bound to the inhibitor W7 determined through isotopically edited and filtered transferred NOEs. This is based on the initial ...Details: Structure of the N-terminal domain of human cardiac troponin C bound to the inhibitor W7 determined through isotopically edited and filtered transferred NOEs. This is based on the initial coordinates of 1LXF, the intraprotein conformational restraints of 1MXL, and target geometries for a calcium-binding loop. The amine moiety of W7 is charged in this structure determination. |
|---|---|
| NMR experiment | Type: 3D-{1H,12C}-filtered-{1H,13C}-edited NOESY |
- Sample preparation
Sample preparation
| Details | Contents: 0.8 mM [U-99% 13C; U-99% 15N] cNTnC, 4.9 mM CALCIUM ION, 0.8 mM N-(6-AMINOHEXYL)-5-CHLORO-1-NAPHTHALENESULFONAMIDE, 10 mM imidazole, 83 mM [U-99% 2H] DSS, 100% D2O Solvent system: 100% D2O | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||
| Sample conditions | pH: 6.75 / Temperature: 303 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: molecular dynamics / Software ordinal: 1 | ||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 50 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller









 PDBj
PDBj





 Xplor-NIH
Xplor-NIH