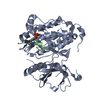
Entry Database : PDB / ID : 2jitTitle Crystal structure of EGFR kinase domain T790M mutation EPIDERMAL GROWTH FACTOR RECEPTOR Keywords / / / / / / / / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species HOMO SAPIENS (human)Method / / / Resolution : 3.1 Å Authors Yun, C.-H. / Mengwasser, K.E. / Toms, A.V. / Woo, M.S. / Greulich, H. / Wong, K.-K. / Meyerson, M. / Eck, M.J. Journal : Proc.Natl.Acad.Sci.USA / Year : 2008Title : The T790M Mutation in Egfr Kinase Causes Drug Resistance by Increasing the Affinity for ATP.Authors : Yun, C.-H. / Mengwasser, K.E. / Toms, A.V. / Woo, M.S. / Greulich, H. / Wong, K. / Meyerson, M. / Eck, M.J. History Deposition Jul 1, 2007 Deposition site / Processing site Revision 1.0 Jan 22, 2008 Provider / Type Revision 1.1 May 7, 2011 Group Revision 1.2 Jul 13, 2011 Group Revision 1.3 Dec 13, 2023 Group Data collection / Database references ... Data collection / Database references / Other / Refinement description Category chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_database_status / pdbx_initial_refinement_model Item / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_sf
Show all Show less Remark 700 SHEET DETERMINATION METHOD: AUTHOR PROVIDED.
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information HOMO SAPIENS (human)
HOMO SAPIENS (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 3.1 Å
MOLECULAR REPLACEMENT / Resolution: 3.1 Å  Authors
Authors Citation
Citation Journal: Proc.Natl.Acad.Sci.USA / Year: 2008
Journal: Proc.Natl.Acad.Sci.USA / Year: 2008 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2jit.cif.gz
2jit.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2jit.ent.gz
pdb2jit.ent.gz PDB format
PDB format 2jit.json.gz
2jit.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/ji/2jit
https://data.pdbj.org/pub/pdb/validation_reports/ji/2jit ftp://data.pdbj.org/pub/pdb/validation_reports/ji/2jit
ftp://data.pdbj.org/pub/pdb/validation_reports/ji/2jit


 Links
Links Assembly
Assembly


 Components
Components HOMO SAPIENS (human) / Description: EGFR 696-1022 T790M / Plasmid: PACG2T / Cell line (production host): SF9 / Production host:
HOMO SAPIENS (human) / Description: EGFR 696-1022 T790M / Plasmid: PACG2T / Cell line (production host): SF9 / Production host: 
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  APS
APS  / Beamline: 19-ID / Wavelength: 0.97924
/ Beamline: 19-ID / Wavelength: 0.97924  Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller





































 PDBj
PDBj












