[English] 日本語
 Yorodumi
Yorodumi- PDB-2hji: Structural model for the Fe-containing isoform of acireductone di... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2hji | ||||||
|---|---|---|---|---|---|---|---|
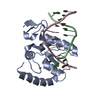
| Title | Structural model for the Fe-containing isoform of acireductone dioxygenase | ||||||
 Components Components | E-2/E-2' protein | ||||||
 Keywords Keywords | OXIDOREDUCTASE / dioxygenase / non-heme iron / isozyme / methionine salvage / structural entropy | ||||||
| Function / homology |  Function and homology information Function and homology informationacireductone dioxygenase (Ni2+-requiring) / acireductone dioxygenase [iron(II)-requiring] / acireductone dioxygenase (Ni2+-requiring) activity / acireductone dioxygenase [iron(II)-requiring] activity / : / : / nickel cation binding / iron ion binding Similarity search - Function | ||||||
| Biological species |  Klebsiella oxytoca (bacteria) Klebsiella oxytoca (bacteria) | ||||||
| Method | SOLUTION NMR / restrained simulated annealing using torsional dynamics | ||||||
 Authors Authors | Pochapsky, T.C. / Ju, T. / Maroney, M.J. / Chai, S.C. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2006 Journal: J.Mol.Biol. / Year: 2006Title: One Protein, Two Enzymes Revisited: A Structural Entropy Switch Interconverts the Two Isoforms of Acireductone Dioxygenase Authors: Ju, T. / Goldsmith, R.B. / Chai, S.C. / Maroney, M.J. / Pochapsky, S.S. / Pochapsky, T.C. #1: Journal: J.Biol.Chem. / Year: 1999 Title: One protein, two enzymes. Authors: Dai, Y. / Wensink, P. / Abeles, R.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2hji.cif.gz 2hji.cif.gz | 674.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2hji.ent.gz pdb2hji.ent.gz | 561 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2hji.json.gz 2hji.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/hj/2hji https://data.pdbj.org/pub/pdb/validation_reports/hj/2hji ftp://data.pdbj.org/pub/pdb/validation_reports/hj/2hji ftp://data.pdbj.org/pub/pdb/validation_reports/hj/2hji | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 20219.412 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Klebsiella oxytoca (bacteria) / Strain: ATCC 8724 / Plasmid: pET3a / Production host: Klebsiella oxytoca (bacteria) / Strain: ATCC 8724 / Plasmid: pET3a / Production host:  References: UniProt: Q9ZFE7, acireductone dioxygenase [iron(II)-requiring] |
|---|---|
| #2: Chemical | ChemComp-FE2 / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||
| NMR details | Text: residual dipolar couplings obtained using filamentous phage (fd) and C12E5 polyol |
- Sample preparation
Sample preparation
| Details | Contents: uniform 15N, 13C labeled, 1 mM / Solvent system: 90/10 H2O/D2O 50 mM KPi buffer, pH 7.5 |
|---|---|
| Sample conditions | Ionic strength: 50 mM KPi / pH: 7.5 / Pressure: 1 atm / Temperature: 298 K |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| NMR spectrometer | Type: Varian UNITYPLUS / Manufacturer: Varian / Model: UNITYPLUS / Field strength: 600 MHz |
- Processing
Processing
| NMR software | Name: CNS / Developer: Brunger A. T. etall / Classification: refinement |
|---|---|
| Refinement | Method: restrained simulated annealing using torsional dynamics Software ordinal: 1 Details: Only residues 1-156 are included in structural models, as residues 157-179 are disordered in solution. |
| NMR representative | Selection criteria: closest to the average |
| NMR ensemble | Conformer selection criteria: back calculated data agree with experimental NOESY spectrum,structures with acceptable covalent geometry Conformers calculated total number: 300 / Conformers submitted total number: 14 |
 Movie
Movie Controller
Controller













 PDBj
PDBj







