+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2fza | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Crystal structure of d(GCGGGAGC): the base-intercalated duplex | ||||||||||||||||||
 Components Components | 5'-D(* Keywords KeywordsDNA / base-intercalated duplex / base-intercalated motif / sheared G:A pair / DNA hexaplex / deoxyribonucleic acid | Function / homology | DNA |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 3.6 Å MOLECULAR REPLACEMENT / Resolution: 3.6 Å  Authors AuthorsKondo, J. / Ciengshin, T. / Juan, E.C.M. / Mitomi, K. / Takenaka, A. |  Citation Citation Journal: Nucleosides Nucleotides Nucleic Acids / Year: 2006 Journal: Nucleosides Nucleotides Nucleic Acids / Year: 2006Title: Crystal structure of d(gcGXGAgc) with X=G: a mutation at X is possible to occur in a base-intercalated duplex for multiplex formation Authors: Kondo, J. / Ciengshin, T. / Juan, E.C.M. / Sato, Y. / Mitomi, K. / Shimizu, S. / Takenaka, A. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2fza.cif.gz 2fza.cif.gz | 19.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2fza.ent.gz pdb2fza.ent.gz | 12 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2fza.json.gz 2fza.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/fz/2fza https://data.pdbj.org/pub/pdb/validation_reports/fz/2fza ftp://data.pdbj.org/pub/pdb/validation_reports/fz/2fza ftp://data.pdbj.org/pub/pdb/validation_reports/fz/2fza | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 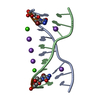
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||||||||||
| Unit cell |
| ||||||||||||||||||
| Components on special symmetry positions |
| ||||||||||||||||||
| Details | The biological assembly is a DNA hexaplex generate from the three DNA duplex (X,Y,Z),(-Y,X-Y,Z) and (-X+Y,-X,Z). |
- Components
Components
| #1: DNA chain | Mass: 2571.538 Da / Num. of mol.: 2 / Source method: obtained synthetically / Details: chemically synthesized #2: Chemical | #3: Chemical | ChemComp-NA / #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.23 Å3/Da / Density % sol: 44.96 % | ||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 7 Details: 50mM sodium cacodylate, 10mM spermine tetrahydrochloride, 20mM calcium chloride, 100mM sodium chloride, 10%(v/v) 2-methyl-2,4-pentanediol, pH 7.0, VAPOR DIFFUSION, HANGING DROP, temperature 293K | ||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
|
-Data collection
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SPring-8 SPring-8  / Beamline: BL44XU / Wavelength: 0.9 Å / Beamline: BL44XU / Wavelength: 0.9 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Type: MAC Science DIP-2040 / Detector: CCD / Date: Jun 3, 2003 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Monochromator: ROTATED-INCLINED SI(111) DOUBLE CRYSTAL MONOCHROMATOR Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.9 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 3.605→31.496 Å / Num. obs: 525 / % possible obs: 93.6 % / Redundancy: 14.5 % / Rmerge(I) obs: 0.112 / Rsym value: 0.112 / Net I/σ(I): 3.1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell |
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 3.6→10 Å / FOM work R set: 0.831 / σ(F): 3 MOLECULAR REPLACEMENT / Resolution: 3.6→10 Å / FOM work R set: 0.831 / σ(F): 3 Stereochemistry target values: G. Parkinson, J. Vojtechovsky, L. Clowney, A.T. Brunger, H.M. Berman, New Parameters for the Refinement of Nucleic Acid Containing Structures, Acta Cryst. D52, 57-64 (1996).
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Bsol: 300 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 3.6→10 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Total num. of bins used: 8
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj






