+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2frg | ||||||
|---|---|---|---|---|---|---|---|
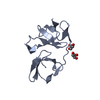
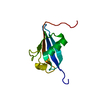
| Title | Structure of the immunoglobulin-like domain of human TLT-1 | ||||||
 Components Components | trem-like transcript-1 | ||||||
 Keywords Keywords | IMMUNE SYSTEM / immunoglobulin-like / beta-sandwich / cell surface receptor / triggering receptor / TREM-like / platelet receptor / ITIM | ||||||
| Function / homology |  Function and homology information Function and homology informationplatelet alpha granule / calcium-mediated signaling / platelet activation / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / transmembrane signaling receptor activity / nuclear speck / innate immune response / cell surface / Golgi apparatus / signal transduction ...platelet alpha granule / calcium-mediated signaling / platelet activation / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / transmembrane signaling receptor activity / nuclear speck / innate immune response / cell surface / Golgi apparatus / signal transduction / membrane / plasma membrane / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.19 Å MOLECULAR REPLACEMENT / Resolution: 1.19 Å | ||||||
 Authors Authors | Gattis, J.L. / Lubkowski, J. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2006 Journal: J.Biol.Chem. / Year: 2006Title: The structure of the extracellular domain of triggering receptor expressed on myeloid cells like transcript-1 and evidence for a naturally occurring soluble fragment. Authors: Gattis, J.L. / Washington, A.V. / Chisholm, M.M. / Quigley, L. / Szyk, A. / McVicar, D.W. / Lubkowski, J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2frg.cif.gz 2frg.cif.gz | 59.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2frg.ent.gz pdb2frg.ent.gz | 42.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2frg.json.gz 2frg.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/fr/2frg https://data.pdbj.org/pub/pdb/validation_reports/fr/2frg ftp://data.pdbj.org/pub/pdb/validation_reports/fr/2frg ftp://data.pdbj.org/pub/pdb/validation_reports/fr/2frg | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| Unit cell |
| |||||||||
| Components on special symmetry positions |
| |||||||||
| Details | Current biochemical and crystallographic evidence suggest that the biologically active form is monomeric. |
- Components
Components
| #1: Antibody | Mass: 11523.272 Da / Num. of mol.: 1 / Fragment: immunoglobulin-like domain 20-125 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: TLT-1 / Plasmid: pET22b / Production host: Homo sapiens (human) / Gene: TLT-1 / Plasmid: pET22b / Production host:  |
|---|---|
| #2: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 2 X-RAY DIFFRACTION / Number of used crystals: 2 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.11 Å3/Da / Density % sol: 41.57 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 5.6 Details: Mother liquor contained 15-19% w/w PEG6000, 0.5M Na/K phosphate pH 5.6. Protein solution contained 10mg/ml TLT-1 in 20mM Tris pH 8. Protein and mother liquor were mixed in equal proportions ...Details: Mother liquor contained 15-19% w/w PEG6000, 0.5M Na/K phosphate pH 5.6. Protein solution contained 10mg/ml TLT-1 in 20mM Tris pH 8. Protein and mother liquor were mixed in equal proportions at 15 C, and hanging drops were suspended over mother liquor., VAPOR DIFFUSION, HANGING DROP, temperature 293K |
-Data collection
| Diffraction |
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source |
| ||||||||||||||||||
| Detector |
| ||||||||||||||||||
| Radiation |
| ||||||||||||||||||
| Radiation wavelength |
| ||||||||||||||||||
| Reflection | Resolution: 1.19→30 Å / Num. all: 32052 / Num. obs: 31487 / % possible obs: 98.2 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 6.4 % / Rsym value: 0.064 / Net I/σ(I): 25.9 | ||||||||||||||||||
| Reflection shell | Resolution: 1.19→1.23 Å / Redundancy: 3.5 % / Mean I/σ(I) obs: 3.2 / Rsym value: 0.31 / % possible all: 87 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: Hybrid model containing structurally conserved residues of TREM-1 (1SMO chain A), NKp44(1HKF), and D1 domain of polymeric immunoglobulin receptor (1XED chain E). Resolution: 1.19→30 Å / Cor.coef. Fo:Fc: 0.957 / Cor.coef. Fo:Fc free: 0.954 / SU B: 1.156 / SU ML: 0.024 / Cross valid method: THROUGHOUT / ESU R: 0.046 / ESU R Free: 0.042 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 13.076 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.144 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.19→30 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.19→1.221 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller














 PDBj
PDBj



