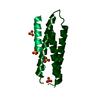
Entry Database : PDB / ID : 2c5jTitle N-terminal domain of tlg1, domain-swapped dimer T-SNARE AFFECTING A LATE GOLGI COMPARTMENT PROTEIN 1 Keywords / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species SACCHAROMYCES CEREVISIAE (brewer's yeast)Method / / Resolution : 2.1 Å Authors Fridmann-Sirkis, Y. / Kent, H.M. / Lewis, M.J. / Evans, P.R. / Pelham, H.R.B. Journal : Traffic / Year : 2006Title : Structural Analysis of the Interaction between the Snare Tlg1 and Vps51.Authors : Fridmann-Sirkis, Y. / Kent, H.M. / Lewis, M.J. / Evans, P.R. / Pelham, H.R.B. History Deposition Oct 27, 2005 Deposition site / Processing site Revision 1.0 Jan 25, 2006 Provider / Type Revision 1.1 May 8, 2011 Group Revision 1.2 Jul 13, 2011 Group Revision 1.3 Dec 13, 2023 Group Data collection / Database references ... Data collection / Database references / Other / Refinement description Category chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_database_status / pdbx_initial_refinement_model Item / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_sf
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.1 Å
MOLECULAR REPLACEMENT / Resolution: 2.1 Å  Authors
Authors Citation
Citation Journal: Traffic / Year: 2006
Journal: Traffic / Year: 2006 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2c5j.cif.gz
2c5j.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2c5j.ent.gz
pdb2c5j.ent.gz PDB format
PDB format 2c5j.json.gz
2c5j.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads 2c5j_validation.pdf.gz
2c5j_validation.pdf.gz wwPDB validaton report
wwPDB validaton report 2c5j_full_validation.pdf.gz
2c5j_full_validation.pdf.gz 2c5j_validation.xml.gz
2c5j_validation.xml.gz 2c5j_validation.cif.gz
2c5j_validation.cif.gz https://data.pdbj.org/pub/pdb/validation_reports/c5/2c5j
https://data.pdbj.org/pub/pdb/validation_reports/c5/2c5j ftp://data.pdbj.org/pub/pdb/validation_reports/c5/2c5j
ftp://data.pdbj.org/pub/pdb/validation_reports/c5/2c5j
 Links
Links Assembly
Assembly
 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418
ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418  Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller
















 PDBj
PDBj
