[English] 日本語
 Yorodumi
Yorodumi- PDB-2bf5: Crystal structure of a toluene 4-monooxygenase catalytic effector... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2bf5 | ||||||
|---|---|---|---|---|---|---|---|
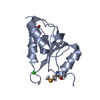
| Title | Crystal structure of a toluene 4-monooxygenase catalytic effector protein variant missing four N-terminal residues (delta-N4 T4moD) | ||||||
 Components Components | TOLUENE-4-MONOOXYGENASE SYSTEM PROTEIN D | ||||||
 Keywords Keywords | OXIDOREDUCTASE / CATALYTIC EFFECTOR PROTEIN / N-TERMINAL TRUNCATED MUTANT / AROMATIC HYDROCARBON CATABOLISM / MONOOXYGENASE / TOLUENE OXIDATION / MOLECULAR REPLACEMENT | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  PSEUDOMONAS MENDOCINA (bacteria) PSEUDOMONAS MENDOCINA (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.71 Å MOLECULAR REPLACEMENT / Resolution: 1.71 Å | ||||||
 Authors Authors | Lountos, G.T. / Mitchell, K.H. / Studts, J.M. / Fox, B.G. / Orville, A.M. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2005 Journal: Biochemistry / Year: 2005Title: Crystal Structures and Functional Studies of T4Mod, the Toluene 4-Monooxygenase Catalytic Effector Protein Authors: Lountos, G.T. / Mitchell, K.H. / Studts, J.M. / Fox, B.G. / Orville, A.M. #1: Journal: Acta Crystallogr.,Sect.D / Year: 2003 Title: Crystallization and Preliminary Analysis of Native and N-Terminal Truncated Isoforms of Toluene-4- Monooxygenase Catalytic Effector Protein Authors: Orville, A.M. / Studts, J.M. / Lountos, G.T. / Mitchell, K.H. / Fox, B.G. #2:  Journal: Biochemistry / Year: 2001 Journal: Biochemistry / Year: 2001Title: Solution Structure of the Toluene 4-Monooxygenase Effector Protein (T4Mod) Authors: Hemmi, H. / Studts, J.M. / Chae, Y.K. / Song, J. / Markley, J.L. / Fox, B.G. #3: Journal: Protein Expr.Purif. / Year: 1999 Title: Application of Fed-Batch Fermentation to the Preparation of Isotopically Labeled or Selenomethionyl-Labeled Proteins Authors: Studts, J.M. / Fox, B.G. #4: Journal: Biochemistry / Year: 2002 Title: Combined Participation of Hydroxylase Active Site Residues and Effector Protein Binding in a Para to Ortho Modulation of Toluene 4-Monooxygenase Regiospecificity Authors: Mitchell, K.H. / Studts, J.M. / Fox, B.G. #5: Journal: Biochemistry / Year: 1996 Title: Recombinant Toluene 4-Monooxygenase: Catalytic and Mossbauer Studies of the Purified Diiron and Rieske Components of a Four Protein Complex Authors: Pikus, J.D. / Studts, J.M. / Achim, C. / Kauffmann, K.E. / Munck, E. / Steffan, R.J. / Mcclay, K. / Fox, B.G. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2bf5.cif.gz 2bf5.cif.gz | 54.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2bf5.ent.gz pdb2bf5.ent.gz | 41.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2bf5.json.gz 2bf5.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bf/2bf5 https://data.pdbj.org/pub/pdb/validation_reports/bf/2bf5 ftp://data.pdbj.org/pub/pdb/validation_reports/bf/2bf5 ftp://data.pdbj.org/pub/pdb/validation_reports/bf/2bf5 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||||
| 2 | 
| ||||||||||||
| Unit cell |
| ||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 11125.419 Da / Num. of mol.: 2 / Fragment: RESIDUES 5-102 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  PSEUDOMONAS MENDOCINA (bacteria) / Strain: KR1 / Plasmid: PET3A / Production host: PSEUDOMONAS MENDOCINA (bacteria) / Strain: KR1 / Plasmid: PET3A / Production host:  References: UniProt: Q00459, Oxidoreductases; Acting on paired donors, with incorporation or reduction of molecular oxygen; With NADH or NADPH as one donor, and incorporation of one atom of oxygen into the other donor #2: Water | ChemComp-HOH / | Sequence details | THE DELTA-N4 T4MOD VARIANT WAS CREATED WITH A VECTOR THAT INITIATES PROTEIN TRANSLATION AT RESIDUE ...THE DELTA-N4 T4MOD VARIANT WAS CREATED WITH A VECTOR THAT INITIATES PROTEIN TRANSLATIO | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.4 Å3/Da / Density % sol: 49 % |
|---|---|
| Crystal grow | Details: PROTEIN WAS CRYSTALLIZED FROM 2.0 M AMMONIUM SULFATE, 5% (V/V) 2-PROPANOL, AND 1.5% (V/V) 1,2,3-HEPTANETRIOL. |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 22-ID / Wavelength: 1.0332 / Beamline: 22-ID / Wavelength: 1.0332 |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Feb 21, 2003 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.0332 Å / Relative weight: 1 |
| Reflection | Resolution: 1.71→37 Å / Num. obs: 23407 / % possible obs: 99.9 % / Observed criterion σ(I): 2 / Redundancy: 21.8 % / Rmerge(I) obs: 0.12 / Net I/σ(I): 3.4 |
| Reflection shell | Resolution: 1.71→1.8 Å / Redundancy: 21.6 % / Rmerge(I) obs: 0.35 / Mean I/σ(I) obs: 1.9 / % possible all: 99.9 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: NATIVE TOLUENE 4-MONOOXYGENASE Resolution: 1.71→37 Å / Cor.coef. Fo:Fc: 0.961 / Cor.coef. Fo:Fc free: 0.945 / SU B: 1.559 / SU ML: 0.053 / Cross valid method: THROUGHOUT / ESU R: 0.095 / ESU R Free: 0.094 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS. RESIDUES 5-10 IN CHAIN A AND CHAIN B AND RESIDUE 102 IN CHAIN B WERE NOT INCLUDED DUE TO THE LACK OF ELECTRON DENSITY.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 13.37 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.71→37 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj