[English] 日本語
 Yorodumi
Yorodumi- PDB-2b3a: Solution structure of the Ras-binding domain of the Ral Guanosine... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2b3a | ||||||
|---|---|---|---|---|---|---|---|
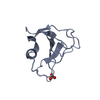
| Title | Solution structure of the Ras-binding domain of the Ral Guanosine Dissociation Stimulator | ||||||
 Components Components | Ral guanine nucleotide dissociation stimulator | ||||||
 Keywords Keywords | SIGNALING PROTEIN / Ras binding domain / ubiquitin fold / signal transduction / automatically solved / AUREMOL | ||||||
| Function / homology |  Function and homology information Function and homology informationGTPase regulator activity / brush border / p38MAPK events / guanyl-nucleotide exchange factor activity / RAF/MAP kinase cascade / Ras protein signal transduction / nucleus / plasma membrane / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method | SOLUTION NMR / simulated annealing first in torsion angle space, simulated annealing in cartesian space. Final refinement in explicit solvent (H2O). | ||||||
 Authors Authors | Gronwald, W. / Maurer, T. / Fuechsl, R. / Wohlgemuth, S. / Herrmann, C. / Kalbitzer, H.R. | ||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: New insights into binding of the possible cancer target RalGDS Authors: Gronwald, W. / Maurer, T. / Fuechsl, R. / Wohlgemuth, S. / Herrmann, C. / Kalbitzer, H.R. #1:  Journal: Nat.Struct.Mol.Biol. / Year: 1997 Journal: Nat.Struct.Mol.Biol. / Year: 1997Title: Structure of the Ras-binding domain of RalGEF and implications for Ras binding and signalling. Authors: Geyer, M. / Herrmann, C. / Wohlgemuth, S. / Wittinghofer, A. / Kalbitzer, H.R. #2:  Journal: Nat.Struct.Mol.Biol. / Year: 1997 Journal: Nat.Struct.Mol.Biol. / Year: 1997Title: Three-dimensional structure of the Ras-interacting domain of RalGDS. Authors: Huang, L. / Weng, X. / Hofer, F. / Martin, G.S. / Kim, S.H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2b3a.cif.gz 2b3a.cif.gz | 277.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2b3a.ent.gz pdb2b3a.ent.gz | 229.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2b3a.json.gz 2b3a.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/b3/2b3a https://data.pdbj.org/pub/pdb/validation_reports/b3/2b3a ftp://data.pdbj.org/pub/pdb/validation_reports/b3/2b3a ftp://data.pdbj.org/pub/pdb/validation_reports/b3/2b3a | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 10052.413 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: RALGDS, RGF / Plasmid: pGEX-4T3 / Species (production host): Escherichia coli / Production host: Homo sapiens (human) / Gene: RALGDS, RGF / Plasmid: pGEX-4T3 / Species (production host): Escherichia coli / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||
| NMR details | Text: sequential assignment bassed on triple resonance spectra, 2D 1H TOCSY, 3D 15N edited TOCSY, 3D 13C edited TOCSY |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions | Ionic strength: 20 mM potassium buffer / pH: 7 / Pressure: ambient / Temperature: 298 K |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Radiation wavelength | Relative weight: 1 | |||||||||||||||
| NMR spectrometer |
|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: simulated annealing first in torsion angle space, simulated annealing in cartesian space. Final refinement in explicit solvent (H2O). Software ordinal: 1 Details: The structures are based on a total of 1680 restraints, 1550 are NOE-derived distance constraints, 104 dihedral angle restraints,26 distance restraints from hydrogen bonds. | ||||||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 300 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller











 PDBj
PDBj



 XPLOR-NIH
XPLOR-NIH