+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: PDB / ID: 2a5r | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
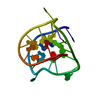
| タイトル | Complex of tetra-(4-n-methylpyridyl) porphin with monomeric parallel-stranded DNA tetraplex, snap-back 3+1 3' G-tetrad, single-residue chain reversal loops, GAG triad in the context of GAAG diagonal loop, C-MYC promoter, NMR, 6 struct. | ||||||||||||||||||||
 要素 要素 | 5'-D(* キーワード キーワードDNA / MONOMERIC PARALLEL-STRANDED QUADRUPLEX / C-MYC PROMOTER 3+1 G-TETRAD / SINGLE NUCLEOTIDE CHAIN REVERSAL LOOP / GAG TRIAD / GAAG LOOP | 機能・相同性 | Chem-POH / DNA / DNA (> 10) |  機能・相同性情報 機能・相同性情報生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト)手法 | 溶液NMR / DISTANCE RESTRAINED MOLECULAR DYNAMICS REFINEMENT |  データ登録者 データ登録者Phan, A.T. / Kuryavyi, V.V. / Gaw, H.Y. / Patel, D.J. |  引用 引用 ジャーナル: Nat.Chem.Biol. / 年: 2005 ジャーナル: Nat.Chem.Biol. / 年: 2005タイトル: Small-molecule interaction with a five-guanine-tract G-quadruplex structure from the human MYC promoter. 著者: Phan, A.T. / Kuryavyi, V. / Gaw, H.Y. / Patel, D.J. 履歴 |
|
- 構造の表示
構造の表示
| 構造ビューア | 分子:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- ダウンロードとリンク
ダウンロードとリンク
- ダウンロード
ダウンロード
| PDBx/mmCIF形式 |  2a5r.cif.gz 2a5r.cif.gz | 108 KB | 表示 |  PDBx/mmCIF形式 PDBx/mmCIF形式 |
|---|---|---|---|---|
| PDB形式 |  pdb2a5r.ent.gz pdb2a5r.ent.gz | 87.9 KB | 表示 |  PDB形式 PDB形式 |
| PDBx/mmJSON形式 |  2a5r.json.gz 2a5r.json.gz | ツリー表示 |  PDBx/mmJSON形式 PDBx/mmJSON形式 | |
| その他 |  その他のダウンロード その他のダウンロード |
-検証レポート
| 文書・要旨 |  2a5r_validation.pdf.gz 2a5r_validation.pdf.gz | 407.1 KB | 表示 |  wwPDB検証レポート wwPDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  2a5r_full_validation.pdf.gz 2a5r_full_validation.pdf.gz | 429.6 KB | 表示 | |
| XML形式データ |  2a5r_validation.xml.gz 2a5r_validation.xml.gz | 2.9 KB | 表示 | |
| CIF形式データ |  2a5r_validation.cif.gz 2a5r_validation.cif.gz | 4.5 KB | 表示 | |
| アーカイブディレクトリ |  https://data.pdbj.org/pub/pdb/validation_reports/a5/2a5r https://data.pdbj.org/pub/pdb/validation_reports/a5/2a5r ftp://data.pdbj.org/pub/pdb/validation_reports/a5/2a5r ftp://data.pdbj.org/pub/pdb/validation_reports/a5/2a5r | HTTPS FTP |
-関連構造データ
- リンク
リンク
- 集合体
集合体
| 登録構造単位 | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR アンサンブル |
|
- 要素
要素
| #1: DNA鎖 | 分子量: 7701.934 Da / 分子数: 1 / 由来タイプ: 天然 / 詳細: C-MYC GENE / 由来: (天然)  Homo sapiens (ヒト) Homo sapiens (ヒト) |
|---|---|
| #2: 化合物 | ChemComp-POH / ( |
-実験情報
-実験
| 実験 | 手法: 溶液NMR | ||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR実験 |
| ||||||||||||||||||||||||||||||||||||||||||||
| NMR実験の詳細 | Text: ONE OF THE EIGHT STRUCTURES OBTAINED FOR FREE DNA TETRAPLEX HAS USED FOR INTERACTIVE MODELING (PDB ID 2A5P). PART OF THE MOLECULE INCLUDING RESIDUES 1,2,3 AND 12 WAS LIFTED AS WHOLE. THE TMPYP4 ...Text: ONE OF THE EIGHT STRUCTURES OBTAINED FOR FREE DNA TETRAPLEX HAS USED FOR INTERACTIVE MODELING (PDB ID 2A5P). PART OF THE MOLECULE INCLUDING RESIDUES 1,2,3 AND 12 WAS LIFTED AS WHOLE. THE TMPYP4 WAS INTERCALATED BETWEEN THE TETRAD G4-G8-G13-G17 AND BASE PAIR A3-A12, GUIDED BY RESTRAINTS BETWEEN THE DRUG AND DNA, AND FAVORABLE POSITIONS OF THE DRUG. BROKEN BONDS BETWEEN THE RESIDUES 11-12-13 AND 3-4 WERE RESTORED, AND MOLECULE WAS SUBJECTED TO MINIMIZATION ROUNDS AND SUBSEQUENT DYNAMICS, WITH THE IMPOSED DNA AND DRUG-DNA RESTRAINTS. INITIALLY, ALL DRUG-DNA RESTRAINTS EXCEPT THESE INVOLVING RESIDUE 1 WERE TREATED AMBIGUOUSLY WITH SUM AVERAGING FROM CONTRIBUTION OF 8 IDENTICAL PROTON ATOMS. THE RESTRAINTS OF THE RESIDUE 1 WERE SPLIT AS ORIGINATING FROM SINGLE (ONE OF 4) PYRIDYL RINGS OF THE TMPYP4 IN FOUR SETS OF COMPUTATIONS. FROM FOUR MOLECULES OBTAINED ONLY ONE WITH LESS VIOLATIONS HAD POSITION OF THE H1' OF THE RESIDUE T1 AS WELL AS H8 OF THE RESIDUE G2 OVER THE AROMATIC RINGS OF THE PORPHYRIN, THUS ACCOUNTING FOR THE UPFIELD SHIFTS OF THESE PROTONS OBSERVED EXPERIMENTALLY. THE POSITION OF THE DRUG IN THE COMPLEX HAS BEEN INCREMENTALLY CHANGED BY ROTATION BY 15 DEG WITHIN THE BOUNDARIES OF VAN DER WAALS SURFACE OF DNA MOLECULE. THE MOLECULE WITH NEW POSITION OF DRUG WAS SUBJECTED TO CONSTRAINED MINIMIZATION AND DYNAMICS RUNS. AFTER THIS ROUND, PYRIDIL RING WAS ALLOWED TO FREE ROTATE AND DYNAMICS AND MINIMIZATION WAS PERFORMED ON SIX COMPLEXES AGAIN. |
- 試料調製
試料調製
| 詳細 |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 試料状態 | イオン強度: 90 mM / pH: 7 / 圧: 1 atm / 温度: 298 K |
-NMR測定
| 放射 | プロトコル: SINGLE WAVELENGTH / 単色(M)・ラウエ(L): M / 散乱光タイプ: x-ray | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 放射波長 | 相対比: 1 | |||||||||||||||
| NMRスペクトロメーター |
|
- 解析
解析
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 精密化 | 手法: DISTANCE RESTRAINED MOLECULAR DYNAMICS REFINEMENT / ソフトェア番号: 1 | ||||||||||||||||||||
| 代表構造 | 選択基準: lowest energy | ||||||||||||||||||||
| NMRアンサンブル | コンフォーマー選択の基準: structures with the least restraint violations 計算したコンフォーマーの数: 6 / 登録したコンフォーマーの数: 6 |
 ムービー
ムービー コントローラー
コントローラー












 PDBj
PDBj







































 HSQC
HSQC