+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1wbf | ||||||
|---|---|---|---|---|---|---|---|
| Title | WINGED BEAN LECTIN, SACCHARIDE FREE FORM | ||||||
 Components Components | PROTEIN (AGGLUTININ) | ||||||
 Keywords Keywords | SUGAR BINDING PROTEIN / LECTIN (AGGLUTININ) / LEGUME LECTIN / PROTEIN CRYSTALLOGRAPHY / BLOOD GROUP SPECIFICITY / SACCHARIDE FREE FORM | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.3 Å MOLECULAR REPLACEMENT / Resolution: 2.3 Å | ||||||
 Authors Authors | Manoj, N. / Srinivas, V.R. / Suguna, K. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.D / Year: 1999 Journal: Acta Crystallogr.,Sect.D / Year: 1999Title: Structure of basic winged-bean lectin and a comparison with its saccharide-bound form. Authors: Manoj, N. / Srinivas, V.R. / Suguna, K. #1:  Journal: J.Mol.Biol. / Year: 1998 Journal: J.Mol.Biol. / Year: 1998Title: Carbohydrate Specificity and Quaternary Association in Basic Winged Bean Lectin: X-Ray Analysis of the Lectin at 2.5 A Resolution Authors: Prabu, M.M. / Sankaranarayanan, R. / Puri, K.D. / Sharma, V. / Surolia, A. / Vijayan, M. / Suguna, K. #2:  Journal: J.Mol.Biol. / Year: 1993 Journal: J.Mol.Biol. / Year: 1993Title: Crystallization and Preliminary X-Ray Studies of the Basic Lectin from Winged Bean (Psophocarpus Tetragonolobus) Authors: Sankaranarayanan, R. / Puri, K.D. / Ganesh, V. / Banerjee, R. / Surolia, A. / Vijayan, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1wbf.cif.gz 1wbf.cif.gz | 112.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1wbf.ent.gz pdb1wbf.ent.gz | 85.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1wbf.json.gz 1wbf.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/wb/1wbf https://data.pdbj.org/pub/pdb/validation_reports/wb/1wbf ftp://data.pdbj.org/pub/pdb/validation_reports/wb/1wbf ftp://data.pdbj.org/pub/pdb/validation_reports/wb/1wbf | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1wblS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 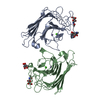
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| Unit cell |
| |||||||||
| Components on special symmetry positions |
| |||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (0.8291, -0.006, 0.559), Vector: |
- Components
Components
| #1: Protein | Mass: 26636.799 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Source: (natural)  #2: Sugar | ChemComp-NAG / #3: Chemical | #4: Chemical | #5: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.26 Å3/Da / Density % sol: 61.1 % | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 7 Details: CRYSTALS WERE GROWN BY BATCH METHOD IN WHICH 25 MICROLITERS OF AN 80 MG/ML PROTEIN SOLUTION IN 0.02M PHOSPHATE BUFFER AT PH 7.0, CONTAINING 0.15M SODIUM CHLORIDE, 0.025 (W/V) SODIUM AZIDE, ...Details: CRYSTALS WERE GROWN BY BATCH METHOD IN WHICH 25 MICROLITERS OF AN 80 MG/ML PROTEIN SOLUTION IN 0.02M PHOSPHATE BUFFER AT PH 7.0, CONTAINING 0.15M SODIUM CHLORIDE, 0.025 (W/V) SODIUM AZIDE, WAS MIXED WITH 45% MPD IN THE SAME BUFFER AT 20 DEGREES CENTIGRADE. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃ / Method: unknown | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 293 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Jun 1, 1995 / Details: MIRRORS |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.3→29.6 Å / Num. obs: 26671 / % possible obs: 88.7 % / Redundancy: 1.5 % / Biso Wilson estimate: 15.4 Å2 / Rmerge(I) obs: 0.073 / Net I/σ(I): 12.5 |
| Reflection shell | Resolution: 2.3→2.4 Å / Redundancy: 1.4 % / Rmerge(I) obs: 0.359 / Mean I/σ(I) obs: 2.8 / % possible all: 80.6 |
| Reflection | *PLUS Num. measured all: 38947 |
| Reflection shell | *PLUS % possible obs: 80.6 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1WBL Resolution: 2.3→8 Å / Rfactor Rfree error: 0.006 / Data cutoff high absF: 10000000 / Data cutoff low absF: 0.001 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 31.6 Å2
| ||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.3→8 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.3→2.44 Å / Rfactor Rfree error: 0.018 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.1 / Classification: refinement X-PLOR / Version: 3.1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller













 PDBj
PDBj








