[English] 日本語
 Yorodumi
Yorodumi- PDB-1uuu: STRUCTURE OF AN RNA HAIRPIN LOOP WITH A 5'-CGUUUCG-3' LOOP MOTIF ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1uuu | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
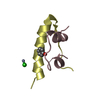
| Title | STRUCTURE OF AN RNA HAIRPIN LOOP WITH A 5'-CGUUUCG-3' LOOP MOTIF BY HETERONUCLEAR NMR SPECTROSCOPY AND DISTANCE GEOMETRY, 15 STRUCTURES | ||||||||||||||||||
 Components Components | RNA (5'-R(* Keywords KeywordsRNA / RIBONUCLEIC ACID / PENTALOOP / HAIRPIN / 18S RIBOSOMAL RNA | Function / homology | RNA / RNA (> 10) |  Function and homology information Function and homology informationMethod | SOLUTION NMR / distance geometry |  Authors AuthorsSich, C. / Ohlenschlager, O. / Ramachandran, R. / Gorlach, M. / Brown, L.R. |  Citation Citation Journal: Biochemistry / Year: 1997 Journal: Biochemistry / Year: 1997Title: Structure of an RNA hairpin loop with a 5'-CGUUUCG-3' loop motif by heteronuclear NMR spectroscopy and distance geometry. Authors: Sich, C. / Ohlenschlager, O. / Ramachandran, R. / Gorlach, M. / Brown, L.R. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1uuu.cif.gz 1uuu.cif.gz | 177.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1uuu.ent.gz pdb1uuu.ent.gz | 148.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1uuu.json.gz 1uuu.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/uu/1uuu https://data.pdbj.org/pub/pdb/validation_reports/uu/1uuu ftp://data.pdbj.org/pub/pdb/validation_reports/uu/1uuu ftp://data.pdbj.org/pub/pdb/validation_reports/uu/1uuu | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: RNA chain | Mass: 6046.608 Da / Num. of mol.: 1 / Source method: obtained synthetically |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
| Sample conditions | pH: 6 / Temperature units: K |
|---|---|
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| Software | Name:  AMBER / Classification: refinement AMBER / Classification: refinement | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR software |
| ||||||||||||
| Refinement | Method: distance geometry / Software ordinal: 1 | ||||||||||||
| NMR ensemble | Conformer selection criteria: LOWEST TARGET FUNCTION / Conformers calculated total number: 300 / Conformers submitted total number: 15 |
 Movie
Movie Controller
Controller











 PDBj
PDBj





























