[English] 日本語
 Yorodumi
Yorodumi- PDB-1tld: CRYSTAL STRUCTURE OF BOVINE BETA-TRYPSIN AT 1.5 ANGSTROMS RESOLUT... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1tld | ||||||
|---|---|---|---|---|---|---|---|
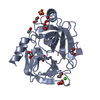
| Title | CRYSTAL STRUCTURE OF BOVINE BETA-TRYPSIN AT 1.5 ANGSTROMS RESOLUTION IN A CRYSTAL FORM WITH LOW MOLECULAR PACKING DENSITY. ACTIVE SITE GEOMETRY, ION PAIRS AND SOLVENT STRUCTURE | ||||||
 Components Components | BETA-TRYPSIN | ||||||
 Keywords Keywords | HYDROLASE / SERINE PROTEINASE | ||||||
| Function / homology |  Function and homology information Function and homology informationtrypsin / serpin family protein binding / serine protease inhibitor complex / digestion / endopeptidase activity / serine-type endopeptidase activity / proteolysis / extracellular space / metal ion binding Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 1.5 Å SYNCHROTRON / Resolution: 1.5 Å | ||||||
 Authors Authors | Bartunik, H.D. / Summers, L.J. / Bartsch, H.H. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1989 Journal: J.Mol.Biol. / Year: 1989Title: Crystal structure of bovine beta-trypsin at 1.5 A resolution in a crystal form with low molecular packing density. Active site geometry, ion pairs and solvent structure. Authors: Bartunik, H.D. / Summers, L.J. / Bartsch, H.H. #1:  Journal: Acta Crystallogr.,Sect.B / Year: 1983 Journal: Acta Crystallogr.,Sect.B / Year: 1983Title: The Geometry of the Reactive Site and of the Peptide Groups in Trypsin, Trypsinogen and its Complexes with Inhibitors Authors: Marquart, M. / Walter, J. / Deisenhofer, J. / Bode, W. / Huber, R. #2:  Journal: Acta Crystallogr.,Sect.B / Year: 1982 Journal: Acta Crystallogr.,Sect.B / Year: 1982Title: On the Disordered Activation Domain in Trypsinogen. Chemical Labelling and Low Temperature Crystallography Authors: Walter, J. / Steigemann, W. / Singh, T.P. / Bartunik, H.D. / Bode, W. / Huber, R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1tld.cif.gz 1tld.cif.gz | 51 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1tld.ent.gz pdb1tld.ent.gz | 36.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1tld.json.gz 1tld.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1tld_validation.pdf.gz 1tld_validation.pdf.gz | 378.5 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1tld_full_validation.pdf.gz 1tld_full_validation.pdf.gz | 381.9 KB | Display | |
| Data in XML |  1tld_validation.xml.gz 1tld_validation.xml.gz | 6.2 KB | Display | |
| Data in CIF |  1tld_validation.cif.gz 1tld_validation.cif.gz | 8.9 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/tl/1tld https://data.pdbj.org/pub/pdb/validation_reports/tl/1tld ftp://data.pdbj.org/pub/pdb/validation_reports/tl/1tld ftp://data.pdbj.org/pub/pdb/validation_reports/tl/1tld | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Atom site foot note | 1: SEE REMARK 4. |
- Components
Components
| #1: Protein | Mass: 23324.287 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  |
|---|---|
| #2: Chemical | ChemComp-CA / |
| #3: Chemical | ChemComp-SO4 / |
| #4: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.99 Å3/Da / Density % sol: 58.81 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Details: THE SOLVENT USED WAS 2.5 M AMMONIUM SULPHATE, 0.1 M PHOSPHATE, 1 MM CACL AT PH 5.3. | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 22 ℃ / pH: 6 / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  EMBL/DESY, HAMBURG EMBL/DESY, HAMBURG  / Beamline: X31 / Wavelength: 1.009 Å / Beamline: X31 / Wavelength: 1.009 Å |
|---|---|
| Detector | Detector: FILM |
| Radiation | Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.009 Å / Relative weight: 1 |
| Reflection | *PLUS Highest resolution: 1.5 Å / Num. obs: 28553 / Num. measured all: 58592 / Rmerge(I) obs: 0.046 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.5→7 Å / Rfactor Rwork: 0.167 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.5→7 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor Rwork: 0.167 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS Type: o_angle_d / Dev ideal: 2.17 |
 Movie
Movie Controller
Controller





















 PDBj
PDBj






