[English] 日本語
 Yorodumi
Yorodumi- PDB-1t5m: Structural transitions as determinants of the action of the calci... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1t5m | ||||||
|---|---|---|---|---|---|---|---|
| Title | Structural transitions as determinants of the action of the calcium-dependent antibiotic daptomycin | ||||||
 Components Components | DAPTOMYCIN | ||||||
 Keywords Keywords | ANTIBIOTIC / DAPTOMYCIN / CUBICIN / LIPOPEPTIDE / CALCIUM-DEPENDENT | ||||||
| Function / homology | Daptomycin / DECANOIC ACID / :  Function and homology information Function and homology information | ||||||
| Biological species |  STREPTOMYCES ROSEOSPORUS (bacteria) STREPTOMYCES ROSEOSPORUS (bacteria) | ||||||
| Method | SOLUTION NMR | ||||||
 Authors Authors | Jung, D. / Rozek, A. / Okon, M. / Hancock, R.E. | ||||||
 Citation Citation |  Journal: Chem.Biol. / Year: 2004 Journal: Chem.Biol. / Year: 2004Title: Structural Transitions as Determinants of the Action of the Calcium-Dependent Antibiotic Daptomycin. Authors: Jung, D. / Rozek, A. / Okon, M. / Hancock, R.E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1t5m.cif.gz 1t5m.cif.gz | 66.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1t5m.ent.gz pdb1t5m.ent.gz | 45.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1t5m.json.gz 1t5m.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/t5/1t5m https://data.pdbj.org/pub/pdb/validation_reports/t5/1t5m ftp://data.pdbj.org/pub/pdb/validation_reports/t5/1t5m ftp://data.pdbj.org/pub/pdb/validation_reports/t5/1t5m | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 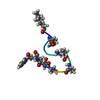
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide |   Type: Cyclic lipopeptide / Class: Antibiotic / Mass: 1484.440 Da / Num. of mol.: 1 / Source method: obtained synthetically Type: Cyclic lipopeptide / Class: Antibiotic / Mass: 1484.440 Da / Num. of mol.: 1 / Source method: obtained syntheticallyDetails: DAPTOMYCIN IS AN ACIDIC CYCLIC LIPOPEPTIDE. THE SCAFFOLD IS MADE OF TWO PARTS: (1) THREE RESIDUES N-TERM EXOCYCLIC PART (2) A DECAPEPTIDE LACTONE RING DERIVED FROM CYCLISATION OF THR3 SIDE ...Details: DAPTOMYCIN IS AN ACIDIC CYCLIC LIPOPEPTIDE. THE SCAFFOLD IS MADE OF TWO PARTS: (1) THREE RESIDUES N-TERM EXOCYCLIC PART (2) A DECAPEPTIDE LACTONE RING DERIVED FROM CYCLISATION OF THR3 SIDE CHAIN ONTO THE C-TER CARBOXYL GROUP THE N-DECANOYL FATTY ACID IS LINKED TO THE MAIN BODY OF THE MOLECULE VIA N-TERM ACYLATION. Source: (synth.)  STREPTOMYCES ROSEOSPORUS (bacteria) / References: NOR: NOR00001, Daptomycin STREPTOMYCES ROSEOSPORUS (bacteria) / References: NOR: NOR00001, Daptomycin |
|---|---|
| #2: Chemical | ChemComp-DKA / 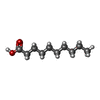  Type: Cyclic lipopeptide / Class: Antibiotic / Mass: 172.265 Da / Num. of mol.: 1 / Source method: obtained synthetically / Formula: C10H20O2 Type: Cyclic lipopeptide / Class: Antibiotic / Mass: 172.265 Da / Num. of mol.: 1 / Source method: obtained synthetically / Formula: C10H20O2Details: DAPTOMYCIN IS AN ACIDIC CYCLIC LIPOPEPTIDE. THE SCAFFOLD IS MADE OF TWO PARTS: (1) THREE RESIDUES N-TERM EXOCYCLIC PART (2) A DECAPEPTIDE LACTONE RING DERIVED FROM CYCLISATION OF THR3 SIDE ...Details: DAPTOMYCIN IS AN ACIDIC CYCLIC LIPOPEPTIDE. THE SCAFFOLD IS MADE OF TWO PARTS: (1) THREE RESIDUES N-TERM EXOCYCLIC PART (2) A DECAPEPTIDE LACTONE RING DERIVED FROM CYCLISATION OF THR3 SIDE CHAIN ONTO THE C-TER CARBOXYL GROUP THE N-DECANOYL FATTY ACID IS LINKED TO THE MAIN BODY OF THE MOLECULE VIA N-TERM ACYLATION. References: Daptomycin |
| Compound details | DAPTOMYCIN IS A CYCLIC TRIDECAMER LIPOPETIDE. HERE, DAPTOMYCIN IS REPRESENTED BY GROUPING TOGETHER ...DAPTOMYCIN |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||
| NMR details | Text: THIS STRUCTURE WAS DETERMINED USING STANDARD 2D HOMONUCLEAR TECHNIQUES. |
- Sample preparation
Sample preparation
| Details | Contents: 2MM DAPTOMYCIN, 100MM KCL, 2MM EGTA, 0.2MM EDTA, 93% H2O, 7% D2O |
|---|---|
| Sample conditions | Ionic strength: 100mM KCL / pH: 6.6 / Pressure: AMBIENT / Temperature: 290.15 K |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| NMR spectrometer | Type: Varian UNITY / Manufacturer: Varian / Model: UNITY / Field strength: 500 MHz |
- Processing
Processing
| NMR software |
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations Conformers calculated total number: 50 / Conformers submitted total number: 15 |
 Movie
Movie Controller
Controller










 PDBj
PDBj