[English] 日本語
 Yorodumi
Yorodumi- PDB-1s2f: Average solution structure of a pseudo-5'-splice site from the ne... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1s2f | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Average solution structure of a pseudo-5'-splice site from the negative regulator of splicing of Rous Sarcoma virus | ||||||||||||||||||||
 Components Components | 5'-R(* Keywords KeywordsRNA / NRS / U1 snRNP binding site / 5' splice site | Function / homology | RNA / RNA (> 10) |  Function and homology information Function and homology informationMethod | SOLUTION NMR / Simulated annealing. Torsion angle dynamics in refinement. | Model type details | minimized average |  Authors AuthorsCabello-Villegas, J. / Giles, K.E. / Soto, A.M. / Yu, P. / Beemon, K.L. / Wang, Y.X. |  Citation Citation Journal: Rna / Year: 2004 Journal: Rna / Year: 2004Title: Solution structure of the pseudo-5' splice site of a retroviral splicing suppressor. Authors: Cabello-Villegas, J. / Giles, K.E. / Soto, A.M. / Yu, P. / Mougin, A. / Beemon, K.L. / Wang, Y.X. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1s2f.cif.gz 1s2f.cif.gz | 23.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1s2f.ent.gz pdb1s2f.ent.gz | 15.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1s2f.json.gz 1s2f.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/s2/1s2f https://data.pdbj.org/pub/pdb/validation_reports/s2/1s2f ftp://data.pdbj.org/pub/pdb/validation_reports/s2/1s2f ftp://data.pdbj.org/pub/pdb/validation_reports/s2/1s2f | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 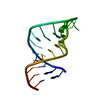
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: RNA chain | Mass: 7350.335 Da / Num. of mol.: 1 / Fragment: Pseudo 5'-splice site / Source method: obtained synthetically / Details: In vitro transcription |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||
| NMR details | Text: Exchangeable proton spectra collected at 15 Celsius |
- Sample preparation
Sample preparation
| Details | Contents: RNA Residues 907-929 from NRS (NRS23) / Solvent system: 10 mM Sodium Phosphate, 25 mM NaCl, pH 6.5 |
|---|---|
| Sample conditions | Ionic strength: 25 mM NaCl, 10 mM NaPi / pH: 6.5 / Pressure: ambient / Temperature: 298 K |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Radiation wavelength | Relative weight: 1 | |||||||||||||||||||||||||
| NMR spectrometer |
|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: Simulated annealing. Torsion angle dynamics in refinement. Software ordinal: 1 Details: 100 structurews were calculated, the minimized average from the 15 lowest energy structures is presented. Constraints used: 423 distance, 63 torsion angle, 117 orientation (RDC). The DELPIC ...Details: 100 structurews were calculated, the minimized average from the 15 lowest energy structures is presented. Constraints used: 423 distance, 63 torsion angle, 117 orientation (RDC). The DELPIC database was employed in the refinement as follows: torsion angle database active for all residues, base-base positional database active for residues 907-912, 914-916, and 921-929. DELPHIC database reference: Clore, G.M. Kuszewski, J. J. Am. Chem. Soc. 125: 1518-1525 | ||||||||||||||||||||
| NMR representative | Selection criteria: minimized average structure | ||||||||||||||||||||
| NMR ensemble | Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller






 PDBj
PDBj





























 Xplor-NIH
Xplor-NIH