[English] 日本語
 Yorodumi
Yorodumi- PDB-1qc0: CRYSTAL STRUCTURE OF A 19 BASE PAIR COPY CONTROL RELATED RNA DUPLEX -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qc0 | ||||||
|---|---|---|---|---|---|---|---|
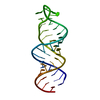
| Title | CRYSTAL STRUCTURE OF A 19 BASE PAIR COPY CONTROL RELATED RNA DUPLEX | ||||||
 Components Components |
| ||||||
 Keywords Keywords | RNA / A-RNA STRUCTURE / RIBONUCLEIC ACID | ||||||
| Function / homology | AMMONIUM ION / RNA / RNA (> 10) Function and homology information Function and homology information | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.55 Å MOLECULAR REPLACEMENT / Resolution: 1.55 Å | ||||||
 Authors Authors | Klosterman, P.S. / Shah, S.A. / Steitz, T.A. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 1999 Journal: Biochemistry / Year: 1999Title: Crystal structures of two plasmid copy control related RNA duplexes: An 18 base pair duplex at 1.20 A resolution and a 19 base pair duplex at 1.55 A resolution. Authors: Klosterman, P.S. / Shah, S.A. / Steitz, T.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qc0.cif.gz 1qc0.cif.gz | 97.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qc0.ent.gz pdb1qc0.ent.gz | 76.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qc0.json.gz 1qc0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qc/1qc0 https://data.pdbj.org/pub/pdb/validation_reports/qc/1qc0 ftp://data.pdbj.org/pub/pdb/validation_reports/qc/1qc0 ftp://data.pdbj.org/pub/pdb/validation_reports/qc/1qc0 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | 
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Unit cell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components on special symmetry positions |
|
- Components
Components
-RNA chain , 4 types, 4 molecules ABCD
| #1: RNA chain | Mass: 2887.767 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: THIS SEQUENCE OCCURS NATURALLY IN E. COLI. |
|---|---|
| #2: RNA chain | Mass: 3135.940 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: THIS SEQUENCE OCCURS NATURALLY IN E. COLI. |
| #3: RNA chain | Mass: 6125.673 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: THIS SEQUENCE OCCURS NATURALLY IN E. COLI. |
| #4: RNA chain | Mass: 6051.681 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: THIS SEQUENCE OCCURS NATURALLY IN E. COLI. |
-Non-polymers , 2 types, 258 molecules 


| #5: Chemical | ChemComp-NH4 / #6: Water | ChemComp-HOH / | |
|---|
-Details
| Compound details | CHAINS A AND B FORM A 9.5 BASE PAIR DUPLEX; A CRYSTALLOGRAPHIC TWO-FOLD AXIS GENERATES THE ...CHAINS A AND B FORM A 9.5 BASE PAIR DUPLEX; A CRYSTALLOG |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 2 X-RAY DIFFRACTION / Number of used crystals: 2 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.3 Å3/Da / Density % sol: 54 % | ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: (NH4)2SO4, SODIUM CACODYLATE, MGCL2, ROP PROTEIN, pH 6.5, VAPOR DIFFUSION, HANGING DROP, temperature 293K | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
| ||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃ / Method: vapor diffusion | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction |
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source |
| ||||||||||||||||||
| Detector |
| ||||||||||||||||||
| Radiation |
| ||||||||||||||||||
| Radiation wavelength |
| ||||||||||||||||||
| Reflection | Resolution: 1.55→40 Å / Num. all: 24898 / Num. obs: 22779 / % possible obs: 91.5 % / Observed criterion σ(F): -3 / Observed criterion σ(I): -3 / Redundancy: 3.6 % / Biso Wilson estimate: 27.2 Å2 / Rmerge(I) obs: 0.052 / Net I/σ(I): 26.9 | ||||||||||||||||||
| Reflection shell | Resolution: 1.55→1.61 Å / Redundancy: 2.7 % / Rmerge(I) obs: 0.452 / Mean I/σ(I) obs: 2.7 / % possible all: 94.8 | ||||||||||||||||||
| Reflection shell | *PLUS % possible obs: 95 % |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: A-RNA Resolution: 1.55→40 Å / Num. parameters: 13059 / Num. restraintsaints: 16977 / Cross valid method: FREE R / σ(F): 0 / σ(I): 0 Stereochemistry target values: PARKINSON, VOJTECHOVSKY, CLOWNEY, BRUNGER & BERMAN, ACTA CRYST D52, 57-64 (1996) Details: ANISOTROPIC REFINEMENT REDUCED FREE R (NO CUTOFF) BY 3.6%
| |||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: MOEWS & KRETSINGER, J.MOL.BIOL. 91, 201-228 (1973) | |||||||||||||||||||||||||||||||||
| Refine analyze | Occupancy sum hydrogen: 561 / Occupancy sum non hydrogen: 1383 | |||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.55→40 Å
| |||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||
| Software | *PLUS Name: SHELXL-97 / Classification: refinement | |||||||||||||||||||||||||||||||||
| Refinement | *PLUS σ(F): 0 / % reflection Rfree: 5 % / Rfactor obs: 0.15 | |||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| |||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Highest resolution: 1.55 Å / Lowest resolution: 1.61 Å / Rfactor Rfree: 0.288 / Rfactor obs: 0.261 |
 Movie
Movie Controller
Controller






 PDBj
PDBj
































