[English] 日本語
 Yorodumi
Yorodumi- PDB-1prw: Crystal structure of bovine brain Ca++ calmodulin in a compact form -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1prw | ||||||
|---|---|---|---|---|---|---|---|
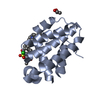
| Title | Crystal structure of bovine brain Ca++ calmodulin in a compact form | ||||||
 Components Components | Calmodulin | ||||||
 Keywords Keywords | METAL BINDING PROTEIN / EF HAND / CALCIUM-BINDING PROTEIN / KINASE ACTIVATOR | ||||||
| Function / homology |  Function and homology information Function and homology informationcalcineurin-mediated signaling / detection of calcium ion / regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / spindle pole / myelin sheath / synaptic vesicle membrane / protein domain specific binding / calcium ion binding / centrosome / protein-containing complex ...calcineurin-mediated signaling / detection of calcium ion / regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / spindle pole / myelin sheath / synaptic vesicle membrane / protein domain specific binding / calcium ion binding / centrosome / protein-containing complex / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SIRSAS / Resolution: 1.7 Å SYNCHROTRON / SIRSAS / Resolution: 1.7 Å | ||||||
 Authors Authors | Fallon, J.L. / Quiocho, F.A. | ||||||
 Citation Citation |  Journal: Structure / Year: 2003 Journal: Structure / Year: 2003Title: A closed compact structure of native ca(2+)-calmodulin. Authors: Fallon, J.L. / Quiocho, F.A. #2:  Journal: Nat.Struct.Biol. / Year: 1995 Journal: Nat.Struct.Biol. / Year: 1995Title: X-ray analysis reveals conformational adaptation of the linker in functional calmodulin mutants Authors: Meador, W.E. / George, S.E. / Means, A.R. / Quiocho, F.A. #3:  Journal: Science / Year: 1993 Journal: Science / Year: 1993Title: Modulation of calmodulin plasticity in molecular recognition on the basis of x-ray structures Authors: Meador, W.E. / Means, A.R. / Quiocho, F.A. #4:  Journal: Science / Year: 1992 Journal: Science / Year: 1992Title: Target enzyme recognition by calmodulin: 2.4 A structure of a calmodulin-peptide complex Authors: Meador, W.E. / Means, A.R. / Quiocho, F.A. #5:  Journal: J.Mol.Biol. / Year: 1992 Journal: J.Mol.Biol. / Year: 1992Title: Calmodulin structure refined at 1.7 A resolution Authors: Chattopadhyaya, R. / Meador, W.E. / Means, A.R. / Quiocho, F.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1prw.cif.gz 1prw.cif.gz | 47.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1prw.ent.gz pdb1prw.ent.gz | 32.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1prw.json.gz 1prw.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pr/1prw https://data.pdbj.org/pub/pdb/validation_reports/pr/1prw ftp://data.pdbj.org/pub/pdb/validation_reports/pr/1prw ftp://data.pdbj.org/pub/pdb/validation_reports/pr/1prw | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 16789.467 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  | ||||
|---|---|---|---|---|---|
| #2: Chemical | ChemComp-CA / #3: Water | ChemComp-HOH / | Has protein modification | Y | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.04 Å3/Da / Density % sol: 39.69 % | ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 296 K / Method: vapor diffusion, hanging drop / pH: 5.4 Details: PEG 6000, sodium acetate, glycerol, pH 5.4, VAPOR DIFFUSION, HANGING DROP, temperature 296K | ||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 23 ℃ / pH: 5.6 / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X4A / Wavelength: 0.9793 Å / Beamline: X4A / Wavelength: 0.9793 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9793 Å / Relative weight: 1 |
| Reflection | Resolution: 1.7→14 Å / Num. all: 15132 / Num. obs: 13383 / % possible obs: 88.4 % / Observed criterion σ(F): 3 / Observed criterion σ(I): 1 / Redundancy: 3.6 % / Biso Wilson estimate: 15.1 Å2 / Rmerge(I) obs: 0.056 / Rsym value: 0.056 / Net I/σ(I): 14.8 |
| Reflection shell | Resolution: 1.7→1.76 Å / Redundancy: 2.96 % / Rmerge(I) obs: 0.38 / Mean I/σ(I) obs: 4.28 / Num. unique all: 1377 / Rsym value: 0.42 / % possible all: 90.7 |
| Reflection | *PLUS Num. obs: 12598 / % possible obs: 92.4 % / Rmerge(I) obs: 0.062 |
| Reflection shell | *PLUS % possible obs: 90.7 % / Rmerge(I) obs: 0.322 / Mean I/σ(I) obs: 4.8 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure: SIRSAS / Resolution: 1.7→14.59 Å / Rfactor Rfree error: 0.006 / Data cutoff high absF: 533665.65 / Data cutoff high rms absF: 533665.65 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 3 / σ(I): 1 / Stereochemistry target values: Engh & Huber Details: The atoms CE, NZ of Lys13 and CD, CE, NZ of Lys21 are present in the file even though the electron density for these atoms is missing
| ||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 83.7254 Å2 / ksol: 0.462227 e/Å3 | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 18.1 Å2
| ||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.7→14.59 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.7→1.81 Å / Rfactor Rfree error: 0.02 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.7 Å / % reflection Rfree: 10 % / Rfactor Rfree: 0.235 / Rfactor Rwork: 0.188 | ||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller














 PDBj
PDBj



