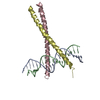
Entry Database : PDB / ID : 1llmTitle Crystal Structure of a Zif23-GCN4 Chimera Bound to DNA 5'-D(*TP*CP*CP*CP*AP*CP*GP*CP*GP*TP*GP*GP*G)-3'chimera of Zif23-GCN4 Keywords / / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Mus musculus (house mouse)Saccharomyces cerevisiae (brewer's yeast)Method / / / Resolution : 1.5 Å Authors Wolfe, S.A. / Grant, R.A. / Pabo, C.O. History Deposition Apr 29, 2002 Deposition site / Processing site Revision 1.0 Sep 30, 2003 Provider / Type Revision 1.1 Apr 28, 2008 Group Revision 1.2 Jul 13, 2011 Group Revision 1.3 Aug 16, 2017 Group / Source and taxonomy / Category / softwareRevision 1.4 Dec 21, 2022 Group / Derived calculationsCategory database_2 / struct_conn ... database_2 / struct_conn / struct_ref_seq_dif / struct_site Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_ref_seq_dif.details / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id Revision 1.5 May 22, 2024 Group / Category / chem_comp_bond
Show all Show less Remark 999 sequence Residues C 151-159 and D 251-259 are linkers between the ZIF23 and GCN4.
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information

 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MIR / Resolution: 1.5 Å
MIR / Resolution: 1.5 Å  Authors
Authors Citation
Citation Journal: Biochemistry / Year: 2003
Journal: Biochemistry / Year: 2003 Journal: Structure / Year: 2000
Journal: Structure / Year: 2000 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 1llm.cif.gz
1llm.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb1llm.ent.gz
pdb1llm.ent.gz PDB format
PDB format 1llm.json.gz
1llm.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/ll/1llm
https://data.pdbj.org/pub/pdb/validation_reports/ll/1llm ftp://data.pdbj.org/pub/pdb/validation_reports/ll/1llm
ftp://data.pdbj.org/pub/pdb/validation_reports/ll/1llm Links
Links Assembly
Assembly
 Components
Components


 X-RAY DIFFRACTION / Number of used crystals: 2
X-RAY DIFFRACTION / Number of used crystals: 2  Sample preparation
Sample preparation Processing
Processing MIR / Resolution: 1.5→20 Å / Rfactor Rfree error: 0.003 / Data cutoff high absF: 10000000 / Data cutoff low absF: 0.001 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 2 / Stereochemistry target values: xplor topology files
MIR / Resolution: 1.5→20 Å / Rfactor Rfree error: 0.003 / Data cutoff high absF: 10000000 / Data cutoff low absF: 0.001 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 2 / Stereochemistry target values: xplor topology files X-PLOR / Classification: refinement
X-PLOR / Classification: refinement Movie
Movie Controller
Controller















 PDBj
PDBj












































