+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1jm0 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CRYSTAL STRUCTURE OF FOUR-HELIX BUNDLE MODEL | |||||||||
 Components Components | PROTEIN (FOUR-HELIX BUNDLE MODEL) | |||||||||
 Keywords Keywords | DE NOVO PROTEIN / ALPHA-HELICAL BUNDLE / PROTEIN DESIGN | |||||||||
| Function / homology | Immunoglobulin FC, subunit C / Single alpha-helices involved in coiled-coils or other helix-helix interfaces / Up-down Bundle / Mainly Alpha / :  Function and homology information Function and homology information | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.7 Å MOLECULAR REPLACEMENT / Resolution: 1.7 Å | |||||||||
 Authors Authors | Di Costanzo, L. / Geremia, S. | |||||||||
 Citation Citation |  Journal: J.Am.Chem.Soc. / Year: 2001 Journal: J.Am.Chem.Soc. / Year: 2001Title: Toward the de novo design of a catalytically active helix bundle: a substrate-accessible carboxylate-bridged dinuclear metal center. Authors: Di Costanzo, L. / Wade, H. / Geremia, S. / Randaccio, L. / Pavone, V. / DeGrado, W.F. / Lombardi, A. #1:  Journal: Proc.Natl.Acad.Sci.USA / Year: 2000 Journal: Proc.Natl.Acad.Sci.USA / Year: 2000Title: Retrostructural Analysis of Metalloproteins: Application to the Design of a Minimal Model for Diiron Proteins Authors: Lombardi, A. / Summa, C.M. / Geremia, S. / Randaccio, L. / Pavone, V. / DeGrado, W.F. #2:  Journal: Curr.Opin.Struct.Biol. / Year: 1999 Journal: Curr.Opin.Struct.Biol. / Year: 1999Title: Tertiary Templates for the Design of Diiron Proteins Authors: Summa, C.M. / Lombardi, A. / Lewis, M. / DeGrado, W.F. #3:  Journal: Annu.Rev.Biochem. / Year: 1999 Journal: Annu.Rev.Biochem. / Year: 1999Title: De Novo Design and Structural Characterization of Proteins and Metalloproteins Authors: DeGrado, W.F. / Summa, C.M. / Pavone, V. / Nastri, F. / Lombardi, A. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1jm0.cif.gz 1jm0.cif.gz | 81.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1jm0.ent.gz pdb1jm0.ent.gz | 62.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1jm0.json.gz 1jm0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jm/1jm0 https://data.pdbj.org/pub/pdb/validation_reports/jm/1jm0 ftp://data.pdbj.org/pub/pdb/validation_reports/jm/1jm0 ftp://data.pdbj.org/pub/pdb/validation_reports/jm/1jm0 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 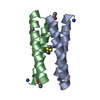
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein/peptide | Mass: 5828.813 Da / Num. of mol.: 6 / Source method: obtained synthetically / Details: THIS PROTEIN WAS CHEMICALLY SYNTHESIZED #2: Chemical | ChemComp-MN / #3: Chemical | #4: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.38 Å3/Da / Density % sol: 42 % | |||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: PEG 400 , Mn(CH3COO)2 , DMSO, Tris-HCl, pH 7.50, VAPOR DIFFUSION, HANGING DROP, temperature 277K | |||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 7.5 | |||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ELETTRA ELETTRA  / Beamline: 5.2R / Wavelength: 1.2 / Beamline: 5.2R / Wavelength: 1.2 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: May 1, 2000 / Details: MIRROR |
| Radiation | Monochromator: SI(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.2 Å / Relative weight: 1 |
| Reflection | Resolution: 1.7→20.1 Å / Num. all: 177455 / Num. obs: 33538 / % possible obs: 99.4 % / Observed criterion σ(F): 0 / Redundancy: 5.3 % / Biso Wilson estimate: 20 Å2 / Rmerge(I) obs: 0.096 / Rsym value: 0.096 / Net I/σ(I): 14 |
| Reflection shell | Resolution: 1.7→1.8 Å / Redundancy: 5.1 % / Rmerge(I) obs: 0.409 / Mean I/σ(I) obs: 2.8 / Rsym value: 0.409 / % possible all: 96.6 |
| Reflection | *PLUS Num. measured all: 177455 |
| Reflection shell | *PLUS % possible obs: 96.6 % / Mean I/σ(I) obs: 2.9 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: THEORETICAL MODEL Resolution: 1.7→20.1 Å / Isotropic thermal model: Isotropic / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 26.02 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.7→20.1 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: REFMAC / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 193.8 Å / Num. reflection obs: 31811 / Rfactor obs: 0.197 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller













 PDBj
PDBj





