[English] 日本語
 Yorodumi
Yorodumi- PDB-1jjx: Solution Structure of Recombinant Human Brain-type Fatty acid Bin... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1jjx | ||||||
|---|---|---|---|---|---|---|---|
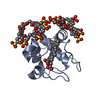
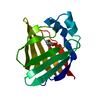
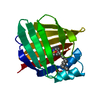
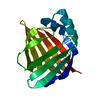
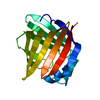
| Title | Solution Structure of Recombinant Human Brain-type Fatty acid Binding Protein | ||||||
 Components Components | BRAIN-TYPE FATTY ACID BINDING PROTEIN | ||||||
 Keywords Keywords | LIPID BINDING PROTEIN / BETA BARREL / FATTY ACID CARRIER / 15N ISOTOPE ENRICHMENT / NMR SPECTROSCOPY | ||||||
| Function / homology |  Function and homology information Function and homology informationNOTCH3 Intracellular Domain Regulates Transcription / Triglyceride catabolism / fatty acid transport / fatty acid binding / epithelial cell proliferation / nervous system development / negative regulation of cell population proliferation / lipid binding / nucleus / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method | SOLUTION NMR / TORSION ANGLE DYNAMICS, SIMULATED ANNEALING, ENERGY MINIMIZATION | ||||||
 Authors Authors | Rademacher, M. / Zimmerman, A.W. / Rueterjans, H. / Veerkamp, J.H. / Luecke, C. | ||||||
 Citation Citation |  Journal: Mol.Cell.Biochem. / Year: 2002 Journal: Mol.Cell.Biochem. / Year: 2002Title: Solution structure of fatty acid-binding protein from human brain. Authors: Rademacher, M. / Zimmerman, A.W. / Ruterjans, H. / Veerkamp, J.H. / Lucke, C. #1:  Journal: J.Biol.Chem. / Year: 2000 Journal: J.Biol.Chem. / Year: 2000Title: Crystal Structure and Thermodynamic Analysis of Human Brain Fatty Acid-Binding Protein Authors: Balendiran, G.K. / Schnuetgen, F. / Scapin, G. / Boerchers, T. / Xhong, N. / Lim, K. / Godbout, R. / Spener, F. / Sacchettini, J.C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1jjx.cif.gz 1jjx.cif.gz | 815.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1jjx.ent.gz pdb1jjx.ent.gz | 683.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1jjx.json.gz 1jjx.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jj/1jjx https://data.pdbj.org/pub/pdb/validation_reports/jj/1jjx ftp://data.pdbj.org/pub/pdb/validation_reports/jj/1jjx ftp://data.pdbj.org/pub/pdb/validation_reports/jj/1jjx | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 14776.702 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Plasmid: PET3D / Species (production host): Escherichia coli / Production host: Homo sapiens (human) / Plasmid: PET3D / Species (production host): Escherichia coli / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 2-3.5MM B-FABP PHOSPHATE BUFFER; 0.05% SODIUM AZIDE |
|---|---|
| Sample conditions | Ionic strength: 20mM POTASSIUM PHOSPHATE / pH: 7 / Pressure: AMBIENT / Temperature: 298.00 K |
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Bruker DMX / Manufacturer: Bruker / Model: DMX / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: TORSION ANGLE DYNAMICS, SIMULATED ANNEALING, ENERGY MINIMIZATION Software ordinal: 1 Details: THE STRUCTURES ARE BASED ON A TOTAL OF 2490 NOE-DERIVED DISTANCE RESTRAINTS | ||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: target function / Conformers calculated total number: 300 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller








 PDBj
PDBj


 HSQC
HSQC