[English] 日本語
 Yorodumi
Yorodumi- PDB-1iii: CRYSTAL STRUCTURE OF THE TRANSTHYRETIN MUTANT TTR Y114C-DATA COLL... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1iii | ||||||
|---|---|---|---|---|---|---|---|
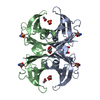
| Title | CRYSTAL STRUCTURE OF THE TRANSTHYRETIN MUTANT TTR Y114C-DATA COLLECTED AT ROOM TEMPERATURE | ||||||
 Components Components | TRANSTHYRETIN | ||||||
 Keywords Keywords | TRANSPORT PROTEIN / GREEK KEY / BETA BARREL | ||||||
| Function / homology |  Function and homology information Function and homology informationDefective visual phototransduction due to STRA6 loss of function / negative regulation of glomerular filtration / The canonical retinoid cycle in rods (twilight vision) / purine nucleobase metabolic process / hormone binding / Non-integrin membrane-ECM interactions / phototransduction, visible light / molecular sequestering activity / retinoid metabolic process / Retinoid metabolism and transport ...Defective visual phototransduction due to STRA6 loss of function / negative regulation of glomerular filtration / The canonical retinoid cycle in rods (twilight vision) / purine nucleobase metabolic process / hormone binding / Non-integrin membrane-ECM interactions / phototransduction, visible light / molecular sequestering activity / retinoid metabolic process / Retinoid metabolism and transport / hormone activity / azurophil granule lumen / Amyloid fiber formation / Neutrophil degranulation / protein-containing complex binding / protein-containing complex / : / extracellular exosome / extracellular region / identical protein binding Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  FOURIER SYNTHESIS / Resolution: 2 Å FOURIER SYNTHESIS / Resolution: 2 Å | ||||||
 Authors Authors | Eneqvist, T. / Olofsson, A. / Ando, Y. / Lundgren, E. / Sauer-Eriksson, A.E. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2002 Journal: Biochemistry / Year: 2002Title: Disulfide-Bond Formation in the Transthyretin Mutant Y114C Prevents Amyloid Fibril Formation in Vivo and in Vitro Authors: Eneqvist, T. / Olofsson, A. / Ando, Y. / Miyakawa, T. / Katsuragi, S. / Jass, J. / Lundgren, E. / Sauer-Eriksson, A.E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1iii.cif.gz 1iii.cif.gz | 60.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1iii.ent.gz pdb1iii.ent.gz | 44.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1iii.json.gz 1iii.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ii/1iii https://data.pdbj.org/pub/pdb/validation_reports/ii/1iii ftp://data.pdbj.org/pub/pdb/validation_reports/ii/1iii ftp://data.pdbj.org/pub/pdb/validation_reports/ii/1iii | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1iikC  1f41S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||
| Unit cell |
| ||||||||||
| Components on special symmetry positions |
| ||||||||||
| Details | The second part of the biological assembly is generated by the two fold axis: -x, -y, z. |
- Components
Components
| #1: Protein | Mass: 13717.329 Da / Num. of mol.: 2 / Fragment: TRANSTHYRETIN / Mutation: Y114C Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: TTR / Plasmid: PET3 / Species (production host): Escherichia coli / Production host: Homo sapiens (human) / Gene: TTR / Plasmid: PET3 / Species (production host): Escherichia coli / Production host:  #2: Chemical | ChemComp-BME / #3: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.24 Å3/Da / Density % sol: 45.04 % | ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 5 Details: 2M ammonium sulphate, 0.1M sodium citrate, 2% PEG 200, 1% BME, pH 5.0, VAPOR DIFFUSION, HANGING DROP at 298K, temperature 298.0K | ||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 7.5 | ||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 298 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: ENRAF-NONIUS / Wavelength: 1.5418 Å ROTATING ANODE / Type: ENRAF-NONIUS / Wavelength: 1.5418 Å |
| Detector | Type: MACSCIENCE / Detector: IMAGE PLATE / Date: Sep 3, 1998 |
| Radiation | Monochromator: Ni FILTER / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2→20 Å / Num. all: 17049 / Num. obs: 17049 / % possible obs: 98.5 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 5.3 % / Biso Wilson estimate: 16.7 Å2 / Rmerge(I) obs: 0.091 / Net I/σ(I): 8.9 |
| Reflection shell | Resolution: 2→2.07 Å / Redundancy: 5.3 % / Rmerge(I) obs: 0.398 / Mean I/σ(I) obs: 4.3 / Num. unique all: 1661 / % possible all: 97.2 |
| Reflection | *PLUS Lowest resolution: 20 Å / Num. measured all: 155823 |
| Reflection shell | *PLUS % possible obs: 97.2 % / Num. unique obs: 1661 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  FOURIER SYNTHESIS FOURIER SYNTHESISStarting model: PDB ENTRY 1F41 Resolution: 2→19.54 Å / SU B: 5.509 / SU ML: 0.1578 / Isotropic thermal model: isotropic / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / ESU R: 0.2109 / ESU R Free: 0.175 / Stereochemistry target values: Engh & Huber Details: Occupancy of BME molecules were refined after structural overall B-factor refinement
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 19.88 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→19.54 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 19.5 Å / Rfactor Rfree: 0.236 / Rfactor Rwork: 0.197 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller























 PDBj
PDBj










