+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1h5b | ||||||
|---|---|---|---|---|---|---|---|
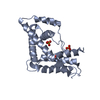
| Title | T cell receptor Valpha11 (AV11S5) domain | ||||||
 Components Components | (MURINE T CELL RECEPTOR (TCR) VALPHA ...) x 3 | ||||||
 Keywords Keywords | T CELL RECEPTOR / IMMUNE RESPONSE / IMMUNOGLOBULIN FOLD | ||||||
| Function / homology | Immunoglobulins / Immunoglobulin-like / Sandwich / Mainly Beta Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 1.85 Å MAD / Resolution: 1.85 Å | ||||||
 Authors Authors | Machius, M. / Cianga, P. / Deisenhofer, J. / Sally Ward, E. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2001 Journal: J.Mol.Biol. / Year: 2001Title: Crystal Structure of a T Cell Receptor Valpha11 (Av11S5) Domain: New Canonical Forms for the First and Second Complementarity Determining Regions Authors: Machius, M. / Cianga, P. / Deisenhofer, J. / Ward, E.S. #1: Journal: J.Mol.Biol. / Year: 2000 Title: Canonical Structures for the Hypervariable Regions of T Cell Ab Receptors Authors: Al-Lazikani, B. / Lesk, A.M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1h5b.cif.gz 1h5b.cif.gz | 109.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1h5b.ent.gz pdb1h5b.ent.gz | 86.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1h5b.json.gz 1h5b.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/h5/1h5b https://data.pdbj.org/pub/pdb/validation_reports/h5/1h5b ftp://data.pdbj.org/pub/pdb/validation_reports/h5/1h5b ftp://data.pdbj.org/pub/pdb/validation_reports/h5/1h5b | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
-MURINE T CELL RECEPTOR (TCR) VALPHA ... , 3 types, 4 molecules ABDC
| #1: Protein | Mass: 12432.696 Da / Num. of mol.: 1 / Fragment: V-ALPHA DOMAIN VA85.33 (AV11S5-AJ17) Source method: isolated from a genetically manipulated source Details: C-TERMINAL HEXA-HISTIDINE TAG / Source: (gene. exp.)   | ||
|---|---|---|---|
| #2: Protein | Mass: 12474.799 Da / Num. of mol.: 2 / Fragment: V-ALPHA DOMAIN VA85.33 (AV11S5-AJ17) Source method: isolated from a genetically manipulated source Details: C-TERMINAL HEXA-HISTIDINE TAG / Source: (gene. exp.)   #3: Protein | | Mass: 12490.799 Da / Num. of mol.: 1 / Fragment: V-ALPHA DOMAIN VA85.33 (AV11S5-AJ17) Source method: isolated from a genetically manipulated source Details: C-TERMINAL HEXA-HISTIDINE TAG / Source: (gene. exp.)   |
-Non-polymers , 3 types, 219 molecules 




| #4: Chemical | | #5: Chemical | ChemComp-GOL / | #6: Water | ChemComp-HOH / | |
|---|
-Details
| Has protein modification | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.7 Å3/Da / Density % sol: 54.1 % / Description: SPOTS HAD SMEARY APPEARANCE AT HIGH RESOLUTION | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Method: vapor diffusion / pH: 5.5 Details: 20C, VAPOR DIFFUSION, 3 UL PROTEIN (5 MG/ML IN 50 MM TRIS/CL, 100 MM NACL, PH 8.0) + 3 UL RESERVOIR SOLUTION (100 MM SODIUM CITRATE/HCL, 1.4-1.7 M LITHIUM CHLORIDE, PH 5.0-6.0) EQUILIBRATED ...Details: 20C, VAPOR DIFFUSION, 3 UL PROTEIN (5 MG/ML IN 50 MM TRIS/CL, 100 MM NACL, PH 8.0) + 3 UL RESERVOIR SOLUTION (100 MM SODIUM CITRATE/HCL, 1.4-1.7 M LITHIUM CHLORIDE, PH 5.0-6.0) EQUILIBRATED AGAINST 1 ML OF RESERVOIR SOLUTION. | ||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃ / pH: 8 / Method: vapor diffusion | ||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 19-ID / Wavelength: 0.9793 / Beamline: 19-ID / Wavelength: 0.9793 |
| Detector | Type: APS1 / Detector: CCD / Date: Feb 15, 1999 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9793 Å / Relative weight: 1 |
| Reflection | Resolution: 1.85→72.34 Å / Num. obs: 45847 / % possible obs: 98.7 % / Observed criterion σ(I): -3 / Redundancy: 6.5 % / Biso Wilson estimate: 22.2 Å2 / Rmerge(I) obs: 0.044 / Net I/σ(I): 30.8 |
| Reflection shell | Resolution: 1.85→1.92 Å / Redundancy: 5.7 % / Rmerge(I) obs: 0.541 / Mean I/σ(I) obs: 3.1 / % possible all: 97.9 |
| Reflection shell | *PLUS % possible obs: 97.9 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MAD / Resolution: 1.85→39.83 Å / Rfactor Rfree error: 0.006 / Data cutoff high absF: 1253020.99 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Details: DISORDERED REGIONS WERE MODELED STEREOCHEMICALLY MAD / Resolution: 1.85→39.83 Å / Rfactor Rfree error: 0.006 / Data cutoff high absF: 1253020.99 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Details: DISORDERED REGIONS WERE MODELED STEREOCHEMICALLY
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 67.5957 Å2 / ksol: 0.385249 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 39.6 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.85→39.83 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.85→1.97 Å / Rfactor Rfree error: 0.017 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Version: 1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj

