+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1h4u | ||||||
|---|---|---|---|---|---|---|---|
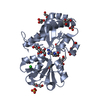
| Title | Domain G2 of mouse nidogen-1 | ||||||
 Components Components | NIDOGEN-1 | ||||||
 Keywords Keywords | EXTRACELLULAR MATRIX PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationLaminin interactions / regulation of basement membrane organization / glomerular basement membrane development / laminin-1 binding / positive regulation of integrin-mediated signaling pathway / Degradation of the extracellular matrix / protein complex involved in cell-matrix adhesion / positive regulation of cell-substrate adhesion / extracellular matrix binding / proteoglycan binding ...Laminin interactions / regulation of basement membrane organization / glomerular basement membrane development / laminin-1 binding / positive regulation of integrin-mediated signaling pathway / Degradation of the extracellular matrix / protein complex involved in cell-matrix adhesion / positive regulation of cell-substrate adhesion / extracellular matrix binding / proteoglycan binding / positive regulation of muscle cell differentiation / basement membrane / collagen binding / laminin binding / extracellular matrix organization / positive regulation of cell adhesion / cell-matrix adhesion / cell periphery / extracellular matrix / calcium ion binding / extracellular region Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MIRAS / Resolution: 2.2 Å MIRAS / Resolution: 2.2 Å | ||||||
 Authors Authors | Hopf, M. / Gohring, W. / Ries, A. / Timpl, R. / Hohenester, E. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 2001 Journal: Nat.Struct.Biol. / Year: 2001Title: Crystal Structure and Mutational Analysis of a Perlecan-Binding Fragment of Nidogen-1 Authors: Hopf, M. / Gohring, W. / Ries, A. / Timpl, R. / Hohenester, E. #1: Journal: Embo J. / Year: 1991 Title: Recombinant Nidogen Consists of Three Globular Domains and Mediates Binding of Laminin to Collagen Type Iv Authors: Fox, J.W. / Mayer, U. / Nischt, R. / Aumailley, M. / Reinhardt, D. / Wiedemann, H. / Mann, K. / Timpl, R. / Krieg, T. / Engel, J. / Chu, M.-L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1h4u.cif.gz 1h4u.cif.gz | 61.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1h4u.ent.gz pdb1h4u.ent.gz | 44.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1h4u.json.gz 1h4u.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/h4/1h4u https://data.pdbj.org/pub/pdb/validation_reports/h4/1h4u ftp://data.pdbj.org/pub/pdb/validation_reports/h4/1h4u ftp://data.pdbj.org/pub/pdb/validation_reports/h4/1h4u | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 29272.756 Da / Num. of mol.: 1 / Fragment: G2 FRAGMENT, RESIDUES 395-659 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   HOMO SAPIENS (human) / References: UniProt: P10493 HOMO SAPIENS (human) / References: UniProt: P10493 |
|---|---|
| #2: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.92 Å3/Da / Density % sol: 57.89 % | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 5.5 Details: 10 MG/ML PROTEIN, 5-8% PEG4000, 0.1 M ACETATE PH 5.5 | |||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 6.8 / Method: vapor diffusion, hanging drop | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SRS SRS  / Beamline: PX9.6 / Wavelength: 0.87 / Beamline: PX9.6 / Wavelength: 0.87 |
| Detector | Type: QUANTUM4 / Detector: CCD / Date: Aug 18, 2000 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.87 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→15 Å / Num. obs: 17203 / % possible obs: 99.7 % / Observed criterion σ(I): 0 / Redundancy: 5 % / Rmerge(I) obs: 0.066 / Net I/σ(I): 14.4 |
| Reflection shell | Resolution: 2.2→2.32 Å / Redundancy: 5 % / Rmerge(I) obs: 0.391 / Mean I/σ(I) obs: 3.8 / % possible all: 100 |
| Reflection | *PLUS Lowest resolution: 15 Å |
| Reflection shell | *PLUS % possible obs: 100 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MIRAS / Resolution: 2.2→15 Å / Data cutoff high absF: 100000 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 MIRAS / Resolution: 2.2→15 Å / Data cutoff high absF: 100000 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 52 Å2 / ksol: 0.33 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→15 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor Rfree: 0.28 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller












 PDBj
PDBj










