[English] 日本語
 Yorodumi
Yorodumi- PDB-1cyc: THE CRYSTAL STRUCTURE OF BONITO (KATSUO) FERROCYTOCHROME C AT 2.3... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1cyc | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | THE CRYSTAL STRUCTURE OF BONITO (KATSUO) FERROCYTOCHROME C AT 2.3 ANGSTROMS RESOLUTION. II. STRUCTURE AND FUNCTION | |||||||||
 Components Components | FERROCYTOCHROME C | |||||||||
 Keywords Keywords | ELECTRON TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationmitochondrial intermembrane space / electron transfer activity / heme binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Katsuwonus pelamis (skipjack tuna) Katsuwonus pelamis (skipjack tuna) | |||||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2.3 Å X-RAY DIFFRACTION / Resolution: 2.3 Å | |||||||||
 Authors Authors | Tanaka, N. / Yamane, T. / Tsukihara, T. / Ashida, T. / Kakudo, M. | |||||||||
 Citation Citation |  Journal: J.Biochem.(Tokyo) / Year: 1975 Journal: J.Biochem.(Tokyo) / Year: 1975Title: The crystal structure of bonito (katsuo) ferrocytochrome c at 2.3 A resolution. II. Structure and function. Authors: Tanaka, N. / Yamane, T. / Tsukihara, T. / Ashida, T. / Kakudo, M. #1:  Journal: J.Biochem.(Tokyo) / Year: 1973 Journal: J.Biochem.(Tokyo) / Year: 1973Title: Oxidation of a Ferrocytochrome C in the Crystalline State-Structural Change and Anion Binding Authors: Tsukihara, T. / Yamane, T. / Tanaka, N. / Ashida, T. / Kakudo, M. #2:  Journal: J.Biochem.(Tokyo) / Year: 1973 Journal: J.Biochem.(Tokyo) / Year: 1973Title: The Crystal Structure of Bonito (Katsuo) Ferrocytochrome C at 2.3 Angstroms Resolution Authors: Ashida, T. / Tanaka, N. / Yamane, T. / Tsukihara, T. / Kakudo, M. #3:  Journal: J.Biochem.(Tokyo) / Year: 1971 Journal: J.Biochem.(Tokyo) / Year: 1971Title: The Crystal Structure of Bonito (Katsuo) Ferrocytochrome C at 4 Angstroms Resolution Authors: Ashida, T. / Ueki, T. / Tsukihara, T. / Sugihara, A. / Takano, T. / Kakudo, M. #4:  Journal: J.Biochem.(Tokyo) / Year: 1971 Journal: J.Biochem.(Tokyo) / Year: 1971Title: The Amino Acid Sequence of Cytochrome C from Bonito (Katsuwonus Pelamis, Linnaeus) Authors: Nakayama, T. / Titani, K. / Narita, K. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1cyc.cif.gz 1cyc.cif.gz | 64.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1cyc.ent.gz pdb1cyc.ent.gz | 30.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1cyc.json.gz 1cyc.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/cy/1cyc https://data.pdbj.org/pub/pdb/validation_reports/cy/1cyc ftp://data.pdbj.org/pub/pdb/validation_reports/cy/1cyc ftp://data.pdbj.org/pub/pdb/validation_reports/cy/1cyc | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 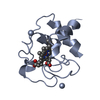
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given / Matrix: (1), |
- Components
Components
| #1: Protein | Mass: 11404.102 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Katsuwonus pelamis (skipjack tuna) / References: UniProt: P00025 Katsuwonus pelamis (skipjack tuna) / References: UniProt: P00025#2: Chemical | #3: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.02 Å3/Da / Density % sol: 39.18 % |
|---|---|
| Crystal grow | *PLUS Method: other / Details: Ashida, T., (1971) J. Biochem., 70, 913. |
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
- Processing
Processing
| Refinement | Highest resolution: 2.3 Å | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement step | Cycle: LAST / Highest resolution: 2.3 Å
|
 Movie
Movie Controller
Controller











 PDBj
PDBj










