[English] 日本語
 Yorodumi
Yorodumi- PDB-1blz: ISOPENICILLIN N SYNTHASE FROM ASPERGILLUS NIDULANS (ACV-FE-NO COMPLEX) -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1blz | ||||||
|---|---|---|---|---|---|---|---|
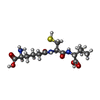
| Title | ISOPENICILLIN N SYNTHASE FROM ASPERGILLUS NIDULANS (ACV-FE-NO COMPLEX) | ||||||
 Components Components | ISOPENICILLIN N SYNTHASE | ||||||
 Keywords Keywords | B-LACTAM ANTIBIOTIC / OXYGENASE / PENICILLIN BIOSYNTHESIS | ||||||
| Function / homology |  Function and homology information Function and homology informationisopenicillin-N synthase / isopenicillin-N synthase activity / penicillin biosynthetic process / L-ascorbic acid binding / iron ion binding / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.45 Å MOLECULAR REPLACEMENT / Resolution: 1.45 Å | ||||||
 Authors Authors | Roach, P.L. / Clifton, I.J. / Hensgens, C.M.H. / Shibata, N. / Schofield, C.J. / Hajdu, J. / Baldwin, J.E. | ||||||
 Citation Citation |  Journal: Nature / Year: 1997 Journal: Nature / Year: 1997Title: Structure of isopenicillin N synthase complexed with substrate and the mechanism of penicillin formation. Authors: Roach, P.L. / Clifton, I.J. / Hensgens, C.M. / Shibata, N. / Schofield, C.J. / Hajdu, J. / Baldwin, J.E. #1:  Journal: Nature / Year: 1995 Journal: Nature / Year: 1995Title: Crystal Structure of Isopenicillin N Synthase is the First from a New Structural Family of Enzymes Authors: Roach, P.L. / Clifton, I.J. / Fulop, V. / Harlos, K. / Barton, G.J. / Hajdu, J. / Andersson, I. / Schofield, C.J. / Baldwin, J.E. #2:  Journal: Protein Sci. / Year: 1995 Journal: Protein Sci. / Year: 1995Title: Crystallization and Preliminary X-Ray Diffraction Studies on Recombinant Isopenicillin N Synthase from Aspergillus Nidulans Authors: Roach, P.L. / Schofield, C.J. / Baldwin, J.E. / Clifton, I.J. / Hajdu, J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1blz.cif.gz 1blz.cif.gz | 82.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1blz.ent.gz pdb1blz.ent.gz | 60.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1blz.json.gz 1blz.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bl/1blz https://data.pdbj.org/pub/pdb/validation_reports/bl/1blz ftp://data.pdbj.org/pub/pdb/validation_reports/bl/1blz ftp://data.pdbj.org/pub/pdb/validation_reports/bl/1blz | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1bk0C  1ipsS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 37563.836 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #2: Chemical | ChemComp-FE / |
| #3: Chemical | ChemComp-ACV / |
| #4: Chemical | ChemComp-NO / |
| #5: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.32 Å3/Da / Density % sol: 47 % | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 8.5 / Details: pH 8.5 | |||||||||||||||
| Crystal grow | *PLUS Temperature: 17 ℃ / Method: vapor diffusion, hanging drop / Details: Roach, P.L., (1996) Eur. J. Biochem., 242, 736. | |||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID2 / Wavelength: 0.99 / Beamline: ID2 / Wavelength: 0.99 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Dec 1, 1996 / Details: TOROIDAL MIRROR |
| Radiation | Monochromator: SI(111) / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.99 Å / Relative weight: 1 |
| Reflection | Resolution: 1.3→25 Å / Num. obs: 80233 / % possible obs: 95.4 % / Redundancy: 3.21 % / Biso Wilson estimate: 22 Å2 / Rmerge(I) obs: 0.055 / Net I/σ(I): 25.9 |
| Reflection shell | Resolution: 1.3→1.35 Å / Redundancy: 3.04 % / Rmerge(I) obs: 0.318 / Mean I/σ(I) obs: 4.1 / % possible all: 93.2 |
| Reflection | *PLUS Highest resolution: 1.45 Å / Lowest resolution: 22 Å / Num. obs: 50924 / % possible obs: 92 % / Num. measured all: 180036 / Rmerge(I) obs: 0.082 |
| Reflection shell | *PLUS Highest resolution: 1.45 Å / Lowest resolution: 1.49 Å / % possible obs: 85.8 % / Num. unique obs: 3298 / Num. measured obs: 10327 / Rmerge(I) obs: 0.519 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1IPS Resolution: 1.45→30 Å / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 21.3 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.45→30 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: REFMAC / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.2054 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller












 PDBj
PDBj