+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1azu | ||||||
|---|---|---|---|---|---|---|---|
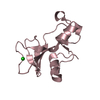
| Title | STRUCTURAL FEATURES OF AZURIN AT 2.7 ANGSTROMS RESOLUTION | ||||||
 Components Components | AZURIN | ||||||
 Keywords Keywords | ELECTRON TRANSPORT (COPPER BINDING) | ||||||
| Function / homology |  Function and homology information Function and homology informationtransition metal ion binding / periplasmic space / electron transfer activity / copper ion binding / zinc ion binding / identical protein binding / plasma membrane Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2.7 Å X-RAY DIFFRACTION / Resolution: 2.7 Å | ||||||
 Authors Authors | Adman, E.T. / Sieker, L.C. / Jensen, L.H. | ||||||
 Citation Citation | Journal: Isr.J.Chem. / Year: 1981 Title: Structural Features of Azurin at 2.7 Angstroms Resolution Authors: Adman, E.T. / Jensen, L.H. #1:  Journal: FEBS Lett. / Year: 1982 Journal: FEBS Lett. / Year: 1982Title: The Effect of Ph and Temperature on the Structure of the Active Site of Azurin from Pseudomonas Aeruginosa Authors: Adman, E.T. / Canters, G.W. / Hill, H.A.O. / Kitchen, N.A. #2:  Journal: Am.Cryst.Assoc.,Abstr.Papers (Summer Meeting) / Year: 1979 Journal: Am.Cryst.Assoc.,Abstr.Papers (Summer Meeting) / Year: 1979Title: Improved Electron Density Maps of Azurin from Pseudomonas Aeruginosa Authors: Adman, E. / Jensen, L.H. #3:  Journal: J.Mol.Biol. / Year: 1978 Journal: J.Mol.Biol. / Year: 1978Title: A Crystallographic Model for Azurin at 3 Angstroms Resolution Authors: Adman, E.T. / Stenkamp, R.E. / Sieker, L.C. / Jensen, L.H. #4:  Journal: Am.Cryst.Assoc.,Abstr.Papers (Summer Meeting) / Year: 1977 Journal: Am.Cryst.Assoc.,Abstr.Papers (Summer Meeting) / Year: 1977Title: A Proposed Model for Azurin at 3 Angstroms Resolution Authors: Adman, E.T. / Stenkamp, R.E. / Sieker, L.C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1azu.cif.gz 1azu.cif.gz | 33.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1azu.ent.gz pdb1azu.ent.gz | 20.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1azu.json.gz 1azu.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/az/1azu https://data.pdbj.org/pub/pdb/validation_reports/az/1azu ftp://data.pdbj.org/pub/pdb/validation_reports/az/1azu ftp://data.pdbj.org/pub/pdb/validation_reports/az/1azu | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 13961.799 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  |
|---|---|
| #2: Chemical | ChemComp-CU / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal grow | *PLUS Method: unknown |
|---|
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
- Processing
Processing
| Software | Name: HENDRICKSON-KONNERT / Version: LEAST-SQUARES REFINEMENT / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Rfactor Rwork: 0.35 / Highest resolution: 2.7 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 2.7 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj

