[English] 日本語
 Yorodumi
Yorodumi- PDB-1abq: CRYSTAL STRUCTURE OF THE UNLIGANDED ABL TYROSINE KINASE SH3 DOMAIN -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1abq | ||||||
|---|---|---|---|---|---|---|---|
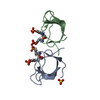
| Title | CRYSTAL STRUCTURE OF THE UNLIGANDED ABL TYROSINE KINASE SH3 DOMAIN | ||||||
 Components Components | ABL TYROSINE KINASE SRC-HOMOLOGY 3 (SH3) DOMAIN | ||||||
 Keywords Keywords | KINASE / SH3 DOMAIN / TRANSFERASE (PHOSPHOTRANSFERASE) / PROTO-ONCOGENE | ||||||
| Function / homology |  Function and homology information Function and homology informationRole of ABL in ROBO-SLIT signaling / HDR through Single Strand Annealing (SSA) / RHO GTPases Activate WASPs and WAVEs / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / Cyclin D associated events in G1 / Turbulent (oscillatory, disturbed) flow shear stress activates signaling by PIEZO1 and integrins in endothelial cells / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / protein localization to cytoplasmic microtubule plus-end / DNA conformation change / DN4 thymocyte differentiation ...Role of ABL in ROBO-SLIT signaling / HDR through Single Strand Annealing (SSA) / RHO GTPases Activate WASPs and WAVEs / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / Cyclin D associated events in G1 / Turbulent (oscillatory, disturbed) flow shear stress activates signaling by PIEZO1 and integrins in endothelial cells / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / protein localization to cytoplasmic microtubule plus-end / DNA conformation change / DN4 thymocyte differentiation / response to epinephrine / phospholipase C-inhibiting G protein-coupled receptor signaling pathway / podocyte apoptotic process / transitional one stage B cell differentiation / regulation of postsynaptic specialization assembly / positive regulation of phospholipase C/protein kinase C signal transduction / regulation of modification of synaptic structure / delta-catenin binding / cerebellum morphogenesis / RUNX1 regulates transcription of genes involved in differentiation of HSCs / regulation of cellular senescence / B cell proliferation involved in immune response / neuroepithelial cell differentiation / positive regulation of extracellular matrix organization / Regulation of actin dynamics for phagocytic cup formation / positive regulation of Wnt signaling pathway, planar cell polarity pathway / microspike assembly / B-1 B cell homeostasis / neuropilin signaling pathway / neuropilin binding / regulation of extracellular matrix organization / Myogenesis / bubble DNA binding / positive regulation of establishment of T cell polarity / activated T cell proliferation / positive regulation of blood vessel branching / proline-rich region binding / negative regulation of mitotic cell cycle / circulatory system development / regulation of Cdc42 protein signal transduction / mitogen-activated protein kinase binding / positive regulation of dendrite development / syntaxin binding / regulation of axon extension / regulation of T cell differentiation / alpha-beta T cell differentiation / positive regulation of cell migration involved in sprouting angiogenesis / negative regulation of cell-cell adhesion / neuromuscular process controlling balance / positive regulation of osteoblast proliferation / platelet-derived growth factor receptor-beta signaling pathway / platelet-derived growth factor receptor signaling pathway / Bergmann glial cell differentiation / cell leading edge / B cell proliferation / negative regulation of cellular senescence / regulation of microtubule polymerization / negative regulation of long-term synaptic potentiation / myoblast proliferation / positive regulation of vasoconstriction / positive regulation of focal adhesion assembly / associative learning / negative regulation of BMP signaling pathway / ephrin receptor signaling pathway / cardiac muscle cell proliferation / negative regulation of endothelial cell apoptotic process / cellular response to transforming growth factor beta stimulus / positive regulation of T cell migration / endothelial cell migration / negative regulation of double-strand break repair via homologous recombination / positive regulation of interleukin-2 production / phagocytosis / ephrin receptor binding / spleen development / four-way junction DNA binding / positive regulation of stress fiber assembly / ruffle / signal transduction in response to DNA damage / phosphotyrosine residue binding / peptidyl-tyrosine phosphorylation / actin filament polymerization / canonical NF-kappaB signal transduction / positive regulation of substrate adhesion-dependent cell spreading / substrate adhesion-dependent cell spreading / positive regulation of mitotic cell cycle / positive regulation of endothelial cell migration / SH2 domain binding / response to endoplasmic reticulum stress / thymus development / protein kinase C binding / positive regulation of release of sequestered calcium ion into cytosol / integrin-mediated signaling pathway / protein serine/threonine kinase activator activity / B cell receptor signaling pathway / post-embryonic development / regulation of actin cytoskeleton organization / non-specific protein-tyrosine kinase / neural tube closure / non-membrane spanning protein tyrosine kinase activity / cell-cell adhesion Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2.8 Å X-RAY DIFFRACTION / Resolution: 2.8 Å | ||||||
 Authors Authors | Musacchio, A. / Wilmanns, M. / Saraste, M. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 1994 Journal: Nat.Struct.Biol. / Year: 1994Title: High-resolution crystal structures of tyrosine kinase SH3 domains complexed with proline-rich peptides. Authors: Musacchio, A. / Saraste, M. / Wilmanns, M. #1:  Journal: Embo J. / Year: 1993 Journal: Embo J. / Year: 1993Title: Crystal Structure of the SH3 Domain in Human Fyn. Comparison of the Three-Dimensional Structures of the SH3 Domain in Tyrosine Kinases and Spectrin Authors: Noble, M.E.M. / Musacchio, A. / Courtneidge, S. / Saraste, M. / Wierenga, R. #2:  Journal: Science / Year: 1993 Journal: Science / Year: 1993Title: Identification of a Ten-Amino Acid SH3 Binding Site Authors: Ren, R. / Mayer, B. / Clark, K.L. / Baltimore, D. #3:  Journal: Nature / Year: 1992 Journal: Nature / Year: 1992Title: Crystal Structure of a Src-Homology 3 (SH3) Domain Authors: Musacchio, A. / Noble, M.E.M. / Pauptit, R. / Wierenga, R. / Saraste, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1abq.cif.gz 1abq.cif.gz | 24.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1abq.ent.gz pdb1abq.ent.gz | 14 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1abq.json.gz 1abq.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ab/1abq https://data.pdbj.org/pub/pdb/validation_reports/ab/1abq ftp://data.pdbj.org/pub/pdb/validation_reports/ab/1abq ftp://data.pdbj.org/pub/pdb/validation_reports/ab/1abq | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 6880.580 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #2: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.55 Å3/Da / Density % sol: 51.73 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS pH: 8.2 / Method: vapor diffusion, hanging drop / Details: used as seeds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction source | Wavelength: 1.5418 Å |
|---|---|
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Sep 12, 1993 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | % possible obs: 96 % / Rmerge(I) obs: 0.08 |
| Reflection | *PLUS Highest resolution: 2.8 Å / Rmerge(I) obs: 0.08 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2.8→8 Å / σ(F): 2 /
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.8→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj







