+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9832 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tetrameric PepTSo2 incorporated in salipro nano particle | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Peptide transporter / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationdipeptide transmembrane transport / tripeptide transmembrane transport / tripeptide transmembrane transporter activity / peptide:proton symporter activity / dipeptide transmembrane transporter activity / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Shewanella oneidensis (bacteria) / Shewanella oneidensis (bacteria) /  Shewanella oneidensis MR-1 (bacteria) Shewanella oneidensis MR-1 (bacteria) | |||||||||
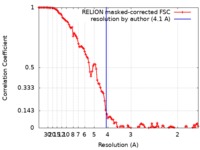
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Kawamoto A / Matoba K | |||||||||
| Funding support |  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: Acta Crystallogr F Struct Biol Commun / Year: 2019 Journal: Acta Crystallogr F Struct Biol Commun / Year: 2019Title: Structural basis for oligomerization of the prokaryotic peptide transporter PepT. Authors: Reina Nagamura / Masahiro Fukuda / Akihiro Kawamoto / Kyoko Matoba / Naoshi Dohmae / Ryuichiro Ishitani / Junichi Takagi / Osamu Nureki /  Abstract: Proton-dependent oligopeptide transporters (POTs) belong to the major facilitator superfamily (MFS) and transport dipeptides and tripeptides from the extracellular environment into the target cell. ...Proton-dependent oligopeptide transporters (POTs) belong to the major facilitator superfamily (MFS) and transport dipeptides and tripeptides from the extracellular environment into the target cell. The human POTs PepT1 and PepT2 are also involved in the absorption of various orally ingested drugs. Previously reported structures revealed that the bacterial POTs possess 14 helices, of which H1-H6 and H7-H12 constitute the typical MFS fold and the residual two helices are involved in the cytoplasmic linker. PepT from Shewanella oneidensis is a unique POT which reportedly assembles as a 200 kDa tetramer. Although the previously reported structures suggested the importance of H12 for tetramer formation, the structural basis for the PepT-specific oligomerization remains unclear owing to the lack of a high-resolution tetrameric structure. In this study, the expression and purification conditions for tetrameric PepT were optimized. A single-particle cryo-EM analysis revealed the tetrameric structure of PepT incorporated into Salipro nanoparticles at 4.1 Å resolution. Furthermore, a combination of lipidic cubic phase (LCP) crystallization and an automated data-processing system for multiple microcrystals enabled crystal structures of PepT to be determined at resolutions of 3.5 and 3.9 Å. The present structures in a lipid bilayer revealed the detailed mechanism for the tetrameric assembly of PepT, in which a characteristic extracellular loop (ECL) interacts with two asparagine residues on H12 which were reported to be important for tetramerization and plays an essential role in oligomeric assembly. This study provides valuable insights into the oligomerization mechanism of this MFS-type transporter, which will further pave the way for understanding other oligomeric membrane proteins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9832.map.gz emd_9832.map.gz | 9.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9832-v30.xml emd-9832-v30.xml emd-9832.xml emd-9832.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_9832_fsc.xml emd_9832_fsc.xml | 10.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_9832.png emd_9832.png | 112.8 KB | ||
| Filedesc metadata |  emd-9832.cif.gz emd-9832.cif.gz | 6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9832 http://ftp.pdbj.org/pub/emdb/structures/EMD-9832 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9832 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9832 | HTTPS FTP |
-Related structure data
| Related structure data |  6ji1MC  6jkcC  6jkdC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9832.map.gz / Format: CCP4 / Size: 107.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9832.map.gz / Format: CCP4 / Size: 107.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.87 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
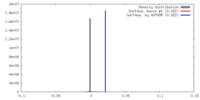
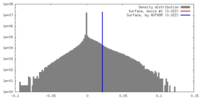
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Tetrameric PepTSo2 with saposin A
| Entire | Name: Tetrameric PepTSo2 with saposin A |
|---|---|
| Components |
|
-Supramolecule #1: Tetrameric PepTSo2 with saposin A
| Supramolecule | Name: Tetrameric PepTSo2 with saposin A / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Shewanella oneidensis (bacteria) Shewanella oneidensis (bacteria) |
-Macromolecule #1: Proton:oligopeptide symporter POT family
| Macromolecule | Name: Proton:oligopeptide symporter POT family / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Shewanella oneidensis MR-1 (bacteria) / Strain: MR-1 Shewanella oneidensis MR-1 (bacteria) / Strain: MR-1 |
| Molecular weight | Theoretical: 57.626559 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTLGTNQVSK THSFMTVSLI ELWERFGYYG MQALIVYFMV QRLGFDDSRA NLVWSACAAL IYVSPAIGGW VGDKILGTKR TMLLGAGIL SVGYALMTVP TENTWFMFSA LGVIVVGNGL FKPNAGNLVR KIYEGDDSKI DSAFTIYYMA VNVGSTFSML L TPWIKDYV ...String: MTLGTNQVSK THSFMTVSLI ELWERFGYYG MQALIVYFMV QRLGFDDSRA NLVWSACAAL IYVSPAIGGW VGDKILGTKR TMLLGAGIL SVGYALMTVP TENTWFMFSA LGVIVVGNGL FKPNAGNLVR KIYEGDDSKI DSAFTIYYMA VNVGSTFSML L TPWIKDYV NAQYGNEFGW HAAFAVCCVG ILVGLGNYAL MHKSLANYGS EPDTRPVNKK SLAIVLALAA LSVVASAIIL EY EDVARVF VYAAGVAVLG IFFHLIRTSE PSERAGLIAA LILTVQTVFF FIFYQQMSTS LALFALRNVD WDFQVFGTHL WTW SPAQFQ ALNPIWIMVL SPVLAWSYSW AGRNNKDFSI AAKFALGFAV VAIGFFIYGF AGQFAVNGKT SSWVMIWGYA SYSL GELLV SGLGLAMIAR YVPARMGGFM MGAYFVASGI SQYLGGVVAN FASVPQDLVD PLQTLPVYTN LFNKLGVAAV VCTII ALAV LPLMRRLTES HHAHSSIENN AAASLRDVKA EQLESSGENL YFQ UniProtKB: Proton:oligopeptide symporter POT family |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blotted for 4.5 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Temperature | Min: 79.55 K / Max: 79.55 K |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number real images: 1609 / Average electron dose: 82.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 96000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-6ji1: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)