+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9747 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Type VI secretion system membrane core complex | |||||||||
 Map data Map data | T6SS membrane core complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Type VI secretion system membrane core complex / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationIcmF-related / Intracellular multiplication and human macrophage-killing / : / : / Type VI secretion protein IcmF helical domain / Type VI secretion system IcmF, C-terminal / Type VI secretion system, lipoprotein SciN / Type VI secretion protein SciN-like superfamily / Type VI secretion protein IcmF C2-like domain / Type VI secretion lipoprotein, VasD, EvfM, TssJ, VC_A0113 / Prokaryotic membrane lipoprotein lipid attachment site profile. Similarity search - Domain/homology | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Meng Y / Zhaofeng Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2019 Journal: Cell Res / Year: 2019Title: Architecture of type VI secretion system membrane core complex. Authors: Meng Yin / Zhaofeng Yan / Xueming Li /  | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9747.map.gz emd_9747.map.gz | 15.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9747-v30.xml emd-9747-v30.xml emd-9747.xml emd-9747.xml | 10.9 KB 10.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9747.png emd_9747.png | 169.9 KB | ||
| Filedesc metadata |  emd-9747.cif.gz emd-9747.cif.gz | 5.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9747 http://ftp.pdbj.org/pub/emdb/structures/EMD-9747 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9747 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9747 | HTTPS FTP |
-Related structure data
| Related structure data |  6ixhMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9747.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9747.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | T6SS membrane core complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
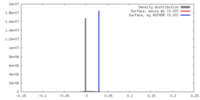
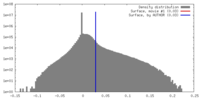
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Type VI Secretion System membrane core complex TssJLM
| Entire | Name: Type VI Secretion System membrane core complex TssJLM |
|---|---|
| Components |
|
-Supramolecule #1: Type VI Secretion System membrane core complex TssJLM
| Supramolecule | Name: Type VI Secretion System membrane core complex TssJLM / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Type VI Secretion System TssJ
| Macromolecule | Name: Type VI Secretion System TssJ / type: protein_or_peptide / ID: 1 / Number of copies: 15 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 19.27109 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAIIAGKAGY GLIIALFSLS LSGCGLTQRV ADGTVSATKS LFYRQIKTLH LDIRAREAIN TSAAGIPLSV VVRIYQLKDN RSFDSADYQ ALFTGDNEIL AGDIIAQKDV WLQPGGSVAV DMPLDDAAKF TGVAAMFLEP DQKKNTWRVV LGRDELEPDT P RLIEVSGN TLTLLPVKDK UniProtKB: Type VI secretion system lipoprotein TssJ |
-Macromolecule #2: Type VI Secretion System TssM
| Macromolecule | Name: Type VI Secretion System TssM / type: protein_or_peptide / ID: 2 / Number of copies: 10 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 127.11307 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNKLACLSGR FGRPGIVFIG VAALWWLITR YGAYLGAETR RDQILLLILL SLGVLFVCYL PVMKKYVQEL TYRRRARKEQ RLPDDEERL AQTPPRYVTV QDIRHTLRRQ YGRFWGRKIR ILLITGTASE VELLTPGLTE QFWQEEQGTL LLWGGDPSQP E NADWLAAL ...String: MNKLACLSGR FGRPGIVFIG VAALWWLITR YGAYLGAETR RDQILLLILL SLGVLFVCYL PVMKKYVQEL TYRRRARKEQ RLPDDEERL AQTPPRYVTV QDIRHTLRRQ YGRFWGRKIR ILLITGTASE VELLTPGLTE QFWQEEQGTL LLWGGDPSQP E NADWLAAL RRLRYRPADG IVWVTSGLSE TLSAPLTEDA LDRVSRAVSS CCERLGWRLP LYVWSLQESP DERGRITQPV GC LLPAECS SDKLKAQLQA MLPGLVAQGI QQICCAPRYY FLLSLAERFR RNIDAVVEPL SVLLRPYRQL LLAGIVFSPA TVG GERSVR HRWRMDNRWE ALPETVQQLP VRLQPSRTGH NWRRSLAVMA AILMMAQGTG MVVSFLANRS LVAEVQEQIR PAQN QQLSP AERLQALLNL QKSLARLQYR EEHGAPWYLR AGMNQNADLL AVVMPLYAQN AHLLLRDAAA AHLEQQLRTF IRLPP DSPQ RGKMAKAAYD QLRLYLMLAQ PQHMEPAWFS RTLMREWPQR DGVSAVFWQA NGPTLLAYYA SGIITHPQWK LTADEE LVS QSRTLLLRHL GTQNSDAMLY QKMLARVAHQ FADMRLTDMT GDTDVSRLFF TDEVVPGMFT RQAWEEAVLP SIDTVIN ER REEMDWVLTD GRQKAPSPVS PEALRQRLTT RYFADFGNAW LNFLNSLHLR KAQTLSDVTE QLTLMADVRQ SPLVALMN T LAVQGCTGQP REAVTDSLVK SARNLLSQEK QPVAVPESRL HGPLATTFGP VLALMDNQNN SADMLNLQTY LTRVTQVRL RLQQIAGSSD PQAMMQLLAQ TVLQGKSVDL TDTRDYGSLT AAGLGQEWYG FGQTVFVRPM EQAWQQVLTP AAESLNARWR TAVVDGWNN AFSGRYPFKN VSSDASLPLL AKYLNTDTGR IARFLQNNLS GVLHREGSRW VPDTINTRGL TFNPAFLKAI N TLSEIADV AFTTGNAGLH FELRPGTAAG VMQTTLITDN QKLIYVNQMP VWKRFTWPAD TEAPGASLSW VSTQAGTRQY AD LPGSWGL IRLLEMARRK AAPGVASGWS LSWQAQDGRM LNYTLRTEAG EGPLVLLKLR NFVLPETVFE LSGTSAFTGN DED AGDTVE ETD UniProtKB: Type VI secretion protein VasK |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 44069 |
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)