[English] 日本語
 Yorodumi
Yorodumi- EMDB-9270: Single-Molecule 3D Image of Cholesteryl Ester Transfer Protein by... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9270 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Single-Molecule 3D Image of Cholesteryl Ester Transfer Protein by Individual-Particle Electron Tomography (No. 01) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
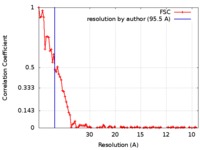
| Method | electron tomography / cryo EM / Resolution: 95.5 Å | |||||||||
 Authors Authors | Wu H / Zhai X / Lei D / Liu J / Yu Y / Bie R / Ren G | |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2018 Journal: Sci Rep / Year: 2018Title: An Algorithm for Enhancing the Image Contrast of Electron Tomography. Authors: Hao Wu / Xiaobo Zhai / Dongsheng Lei / Jianfang Liu / Yadong Yu / Rongfang Bie / Gang Ren /   Abstract: Three-dimensional (3D) reconstruction of a single protein molecule is essential for understanding the relationship between the structural dynamics and functions of the protein. Electron tomography ...Three-dimensional (3D) reconstruction of a single protein molecule is essential for understanding the relationship between the structural dynamics and functions of the protein. Electron tomography (ET) provides a tool for imaging an individual particle of protein from a series of tilted angles. Individual-particle electron tomography (IPET) provides an approach for reconstructing a 3D density map from a single targeted protein particle (without averaging from different particles of this type of protein), in which the target particle was imaged from a series of tilting angles. However, owing to radiation damage limitations, low-dose images (high noise, and low image contrast) are often challenging to be aligned for 3D reconstruction at intermediate resolution (1-3 nm). Here, we propose a computational method to enhance the image contrast, without increasing any experimental dose, for IPET 3D reconstruction. Using an edge-preserving smoothing-based multi-scale image decomposition algorithm, this method can detect the object against a high-noise background and enhance the object image contrast without increasing the noise level or significantly decreasing the image resolution. The method was validated by using both negative staining (NS) ET and cryo-ET images. The successful 3D reconstruction of a small molecule (<100 kDa) indicated that this method can be used as a supporting tool to current ET 3D reconstruction methods for studying protein dynamics via structure determination from each individual particle of the same type of protein. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9270.map.gz emd_9270.map.gz | 789.7 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9270-v30.xml emd-9270-v30.xml emd-9270.xml emd-9270.xml | 9.9 KB 9.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_9270_fsc.xml emd_9270_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_9270.png emd_9270.png | 10.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9270 http://ftp.pdbj.org/pub/emdb/structures/EMD-9270 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9270 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9270 | HTTPS FTP |
-Related structure data
| Related structure data |  9262C  9263C  9266C  9267C  9268C  9269C  9271C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_9270.map.gz / Format: CCP4 / Size: 844.7 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9270.map.gz / Format: CCP4 / Size: 844.7 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.8 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
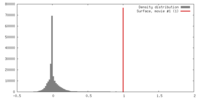
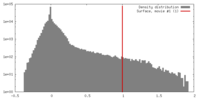
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Cholesteryl ester transfer protein
| Entire | Name: Cholesteryl ester transfer protein |
|---|---|
| Components |
|
-Supramolecule #1: Cholesteryl ester transfer protein
| Supramolecule | Name: Cholesteryl ester transfer protein / type: organelle_or_cellular_component / ID: 1 / Parent: 0 Details: Cholesteryl ester transfer protein (CETP) was produced by MERCK. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Tissue: Blood Homo sapiens (human) / Tissue: Blood |
| Molecular weight | Theoretical: 2.5 MDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 / Details: 1X Dulbeccos phosphate-buffered saline |
|---|---|
| Grid | Model: Homemade / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: LEICA EM GP |
| Details | Cholesteryl ester transfer protein (CETP) was produced by MERCK. |
| Sectioning | Other: NO SECTIONING |
- Electron microscopy
Electron microscopy
| Microscope | ZEISS LIBRA120PLUS |
|---|---|
| Specialist optics | Energy filter - Name: In-column Omega Filter / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Average electron dose: 0.32 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.2 mm / Nominal magnification: 50000 |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)