+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8986 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | A 3.1 Angstrom sub-tomogram average of HIV-1 CA-SP1. | |||||||||
 Map data Map data | Map with a B-factor of 150 applied, as used in generating figures. | |||||||||
 Sample Sample |
| |||||||||
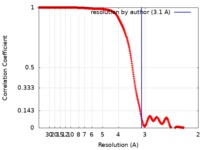
| Method | subtomogram averaging / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Himes BA / Zhang P | |||||||||
 Citation Citation |  Journal: Nat Methods / Year: 2018 Journal: Nat Methods / Year: 2018Title: emClarity: software for high-resolution cryo-electron tomography and subtomogram averaging. Authors: Benjamin A Himes / Peijun Zhang /   Abstract: Macromolecular complexes are intrinsically flexible and often challenging to purify for structure determination by single-particle cryo-electron microscopy (cryo-EM). Such complexes can be studied by ...Macromolecular complexes are intrinsically flexible and often challenging to purify for structure determination by single-particle cryo-electron microscopy (cryo-EM). Such complexes can be studied by cryo-electron tomography (cryo-ET) combined with subtomogram alignment and classification, which in exceptional cases achieves subnanometer resolution, yielding insight into structure-function relationships. However, it remains challenging to apply this approach to specimens that exhibit conformational or compositional heterogeneity or are present in low abundance. To address this, we developed emClarity ( https://github.com/bHimes/emClarity/wiki ), a GPU-accelerated image-processing package featuring an iterative tomographic tilt-series refinement algorithm that uses subtomograms as fiducial markers and a 3D-sampling-function-compensated, multi-scale principal component analysis classification method. We demonstrate that our approach offers substantial improvement in the resolution of maps and in the separation of different functional states of macromolecular complexes compared with current state-of-the-art software. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8986.map.gz emd_8986.map.gz | 15.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8986-v30.xml emd-8986-v30.xml emd-8986.xml emd-8986.xml | 10.1 KB 10.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8986_fsc.xml emd_8986_fsc.xml | 34.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_8986.png emd_8986.png | 46.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8986 http://ftp.pdbj.org/pub/emdb/structures/EMD-8986 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8986 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8986 | HTTPS FTP |
-Related structure data
| Related structure data |  8799C  8802C  8803C  8804C  8805C  8806C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8986.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8986.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map with a B-factor of 150 applied, as used in generating figures. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
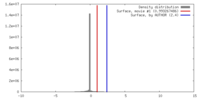
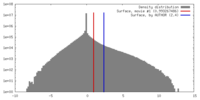
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : HIV-1 CA-SP1
| Entire | Name: HIV-1 CA-SP1 |
|---|---|
| Components |
|
-Supramolecule #1: HIV-1 CA-SP1
| Supramolecule | Name: HIV-1 CA-SP1 / type: virus / ID: 1 / Parent: 0 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: OTHER / Virus enveloped: No / Virus empty: Yes |
|---|
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
| Details | Please refer to EMD-3782/ EMPIAR-10164 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Details | Please refer to EMD-3782/ EMPIAR-10164 |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 3.4 e/Å2 / Details: Please refer to EMD-3782/ EMPIAR-10164 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)