+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8926 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mutated p53-DNA assembly | |||||||||
 Map data Map data | p53 tetramer assembly | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  Human (human) Human (human) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 15.5 Å | |||||||||
 Authors Authors | Kelly DF / Dearnaley WJ / Liang Y | |||||||||
 Citation Citation |  Journal: DNA Repair (Amst) / Year: 2019 Journal: DNA Repair (Amst) / Year: 2019Title: Correcting errors in the BRCA1 warning system. Authors: Yanping Liang / William J Dearnaley / Nick A Alden / Maria J Solares / Brian L Gilmore / Kevin J Pridham / A Cameron Varano / Zhi Sheng / Elizabeth Alli / Deborah F Kelly /  Abstract: Given its important role in human health and disease, remarkably little is known about the full-length three-dimensional (3D) molecular architecture of the breast cancer type 1 susceptibility protein ...Given its important role in human health and disease, remarkably little is known about the full-length three-dimensional (3D) molecular architecture of the breast cancer type 1 susceptibility protein (BRCA1), or its mechanisms to engage the tumor suppressor, TP53 (p53). Here, we show how a prevalent cancer-related mutation in the C-terminal region of the full-length protein, BRCA1, affects its structural properties, yet can be biochemically corrected to restore its functional capacity. As a downstream consequence of restoring the ubiquitin ligase activity of mutated BRCA1, the DNA repair response of p53 was enhanced in cellular extracts naturally deficient in BRCA1 protein expression. Complementary structural insights of p53 tetramers bound to DNA in different stage of the repair process support these biochemical findings in the context of human cancer cells. Equally important, we show how this knowledge can be used to lower the viability of breast cancer cells by modulating the stability of the BRCA1 protein and its associated players. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8926.map.gz emd_8926.map.gz | 112.2 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8926-v30.xml emd-8926-v30.xml emd-8926.xml emd-8926.xml | 12.6 KB 12.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8926.png emd_8926.png | 86.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8926 http://ftp.pdbj.org/pub/emdb/structures/EMD-8926 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8926 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8926 | HTTPS FTP |
-Validation report
| Summary document |  emd_8926_validation.pdf.gz emd_8926_validation.pdf.gz | 78.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_8926_full_validation.pdf.gz emd_8926_full_validation.pdf.gz | 77.7 KB | Display | |
| Data in XML |  emd_8926_validation.xml.gz emd_8926_validation.xml.gz | 495 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8926 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8926 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8926 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8926 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8926.map.gz / Format: CCP4 / Size: 3.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8926.map.gz / Format: CCP4 / Size: 3.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | p53 tetramer assembly | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.4 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
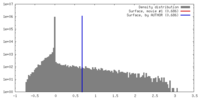
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Mutated p53-DNA assembly
| Entire | Name: Mutated p53-DNA assembly |
|---|---|
| Components |
|
-Supramolecule #1: Mutated p53-DNA assembly
| Supramolecule | Name: Mutated p53-DNA assembly / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Tissue: Breast Homo sapiens (human) / Tissue: Breast |
| Molecular weight | Theoretical: 220 KDa |
-Macromolecule #1: Mutated p53-DNA assembly
| Macromolecule | Name: Mutated p53-DNA assembly / type: protein_or_peptide / ID: 1 / Enantiomer: DEXTRO |
|---|---|
| Source (natural) | Organism:  Human (human) / Tissue: Breast Human (human) / Tissue: Breast |
| Sequence | String: MEEPQSDPSV EPPLSQETFS DLWKLLPENN VLSPLPSQAM DDLMLSPDDI EQWFTEDPGP DEAPRMPEAA PPVAPAPAAP TPAAPAPAPS WPLSSSVPSQ KTYQGSYGFR LGFLHSGTAK SVTCTYSPAL NKMFCQLAKT CPVQLWVDST PPPGTRVRAM AIYKQSQHMT ...String: MEEPQSDPSV EPPLSQETFS DLWKLLPENN VLSPLPSQAM DDLMLSPDDI EQWFTEDPGP DEAPRMPEAA PPVAPAPAAP TPAAPAPAPS WPLSSSVPSQ KTYQGSYGFR LGFLHSGTAK SVTCTYSPAL NKMFCQLAKT CPVQLWVDST PPPGTRVRAM AIYKQSQHMT EVVRRCPHHE RCSDSDGLAP PQHLIRVEGN LRVEYLDDRN TFRHSVVVPY EPPEVGSDCT TIHYNYMCNS SCMGGMNRRP ILTIITLEDS SGNLLGRNSF EVRVCACPGR DRRTEEENLR KKGEPHHELP PGSTKR |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.02 mg/mL |
|---|---|
| Buffer | pH: 7.2 / Component - Name: HEPES buffer Details: 20 mM HEPES, pH 7.2, 150 mM NaCl, 10 mM CaCl2, 10 mM MgCl2 |
| Staining | Type: NEGATIVE / Material: Uranyl Formate / Details: 1% uranyl formate |
| Grid | Material: COPPER / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.026 kPa Details: Grid was coated with amyl acetate followed by a layer of carbon prior to glow discharge. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI SPIRIT |
|---|---|
| Image recording | Film or detector model: FEI EAGLE (2k x 2k) / Digitization - Sampling interval: 30.0 µm / Number grids imaged: 1 / Number real images: 92 / Average electron dose: 5.0 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus min: -1.5 µm / Nominal magnification: 68000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Tecnai Spirit / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)